Polymeric Nanoparticles vs. Liposomes: A Detailed Comparison of Structure, Composition, and Drug Delivery Applications
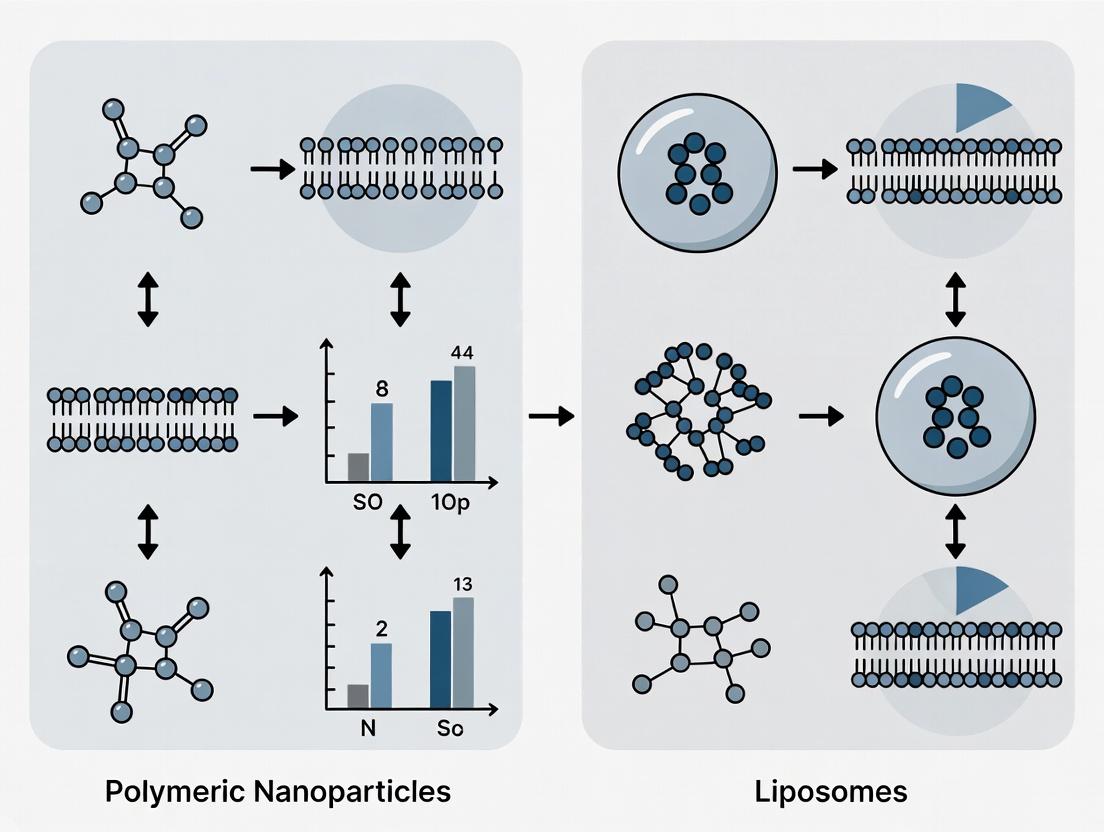
This article provides a comprehensive analysis of polymeric nanoparticles and liposomes, two leading nanocarrier systems in drug delivery.
Polymeric Nanoparticles vs. Liposomes: A Detailed Comparison of Structure, Composition, and Drug Delivery Applications
Abstract
This article provides a comprehensive analysis of polymeric nanoparticles and liposomes, two leading nanocarrier systems in drug delivery. Aimed at researchers, scientists, and drug development professionals, it explores their fundamental structural and compositional differences, core synthesis methods, and key applications. The content delves into critical challenges, scale-up considerations, and optimization strategies for each platform. Finally, a direct comparative evaluation examines stability, biocompatibility, drug loading efficiency, and regulatory progress, offering actionable insights for selecting the optimal nanocarrier for specific therapeutic goals.
Core Blueprints: Deconstructing the Structural and Chemical Foundations of PNPs and Liposomes
Within the context of drug delivery research, the fundamental structure and composition of a nanocarrier dictate its fate, function, and efficacy. Two of the most extensively studied platforms are polymeric nanoparticles (PNPs) and liposomes. This technical guide provides an in-depth comparison of their core definitions, compositions, synthesis, and characterization, serving as a foundation for rational platform selection in advanced therapeutic development.
Core Definitions and Structural Composition
Polymeric Nanoparticles (PNPs) are solid colloidal particles typically ranging from 10 to 1000 nm. They consist of biodegradable or biocompatible polymers, forming a dense matrix where the active ingredient can be dissolved, entrapped, encapsulated, or chemically attached. The core structure is defined by the polymer chain arrangement, which can be configured as a nanosphere (matrix system) or a nanocapsule (reservoir system with a polymer shell surrounding an oily or aqueous core).
Liposomes are spherical vesicles composed of one or more concentric phospholipid bilayers enclosing an aqueous compartment. Their structure mimics a biological membrane, forming a natural barrier. Based on lamellarity and size, they are classified as Small Unilamellar Vesicles (SUVs, 20-100 nm), Large Unilamellar Vesicles (LUVs, 100-1000 nm), or Multilamellar Vesicles (MLVs, >500 nm).
Synthesis and Fabrication: Key Methodologies
Polymeric Nanoparticle Preparation
Method: Nanoprecipitation (Solvent Displacement)
- Principle: A water-miscible organic solvent containing the polymer and hydrophobic drug is added to an aqueous phase with a stabilizer (e.g., PVA, poloxamer). Rapid solvent diffusion leads to polymer precipitation into nanoparticles.
- Detailed Protocol:
- Dissolve 50-200 mg of polymer (e.g., PLGA, PLA) and 5-20 mg of active pharmaceutical ingredient (API) in 10-20 mL of acetone or acetonitrile (organic phase).
- Prepare an aqueous phase containing 1-2% w/v stabilizer (e.g., PVA, polysorbate 80) in 50 mL of deionized water.
- Inject the organic phase into the aqueous phase under moderate magnetic stirring (500-800 rpm) at room temperature.
- Stir for 3-6 hours to allow complete solvent evaporation and nanoparticle hardening.
- Purify by centrifugation (e.g., 20,000 x g, 30 min, 4°C) and resuspend in buffer or lyophilize for storage.
Method: Emulsion-Solvent Evaporation (for hydrophobic drugs)
- Principle: An oil-in-water (O/W) emulsion is formed where the organic solvent (e.g., dichloromethane) containing polymer/drug is emulsified in water. Solvent evaporation solidifies the droplets into nanoparticles.
- Detailed Protocol:
- Dissolve polymer and drug in a volatile organic solvent (e.g., dichloromethane, ethyl acetate).
- Emulsify this organic phase in an aqueous surfactant solution using high-speed homogenization (e.g., 10,000 rpm for 2 min) or probe sonication (e.g., 70% amplitude, 30 s pulses).
- The resulting emulsion is stirred overnight to evaporate the organic solvent.
- Centrifuge and wash as described above.
Liposome Preparation
Method: Thin-Film Hydration & Extrusion
- Principle: Phospholipids are dissolved in an organic solvent, which is evaporated to form a thin lipid film. The film is hydrated with an aqueous buffer, spontaneously forming multilamellar vesicles (MLVs). These are then sized down to unilamellar vesicles via extrusion.
- Detailed Protocol:
- Dissolve phospholipids (e.g., DPPC, DSPC, cholesterol at a molar ratio of 55:45) and any lipid-conjugated PEG (e.g., DSPE-PEG2000) in chloroform in a round-bottom flask.
- Remove the solvent using a rotary evaporator under reduced pressure (e.g., 200 mbar, 40°C water bath) to form a thin, dry lipid film.
- Hydrate the film with an appropriate aqueous buffer (e.g., PBS, HEPES, possibly containing a hydrophilic drug) above the lipid phase transition temperature (e.g., 55°C for DPPC) with gentle agitation for 1 hour.
- Subject the resulting MLV suspension to 5 freeze-thaw cycles (liquid nitrogen/50°C water bath).
- Extrude the suspension 11-21 times through polycarbonate membranes of defined pore size (e.g., 100 nm, then 50 nm) using a thermobarrel extruder set above the lipid transition temperature.
- Purify via dialysis or size-exclusion chromatography to remove unencapsulated material.
Comparative Analysis: Quantitative Data
Table 1: Core Structural & Compositional Comparison
| Parameter | Polymeric Nanoparticles (PNPs) | Liposomes |
|---|---|---|
| Core Structure | Solid polymer matrix or reservoir. | Aqueous core enclosed by lipid bilayer(s). |
| Typical Size Range | 50-500 nm. | 50-200 nm (for SUV/LUV used in delivery). |
| Key Composition | Biodegradable polymers (PLGA, PLA, chitosan, polycaprolactone). | Phospholipids (phosphatidylcholines), cholesterol, PEG-lipids. |
| Drug Loading | Incorporation into matrix/adsorption/conjugation. High for hydrophobic drugs. | Encapsulation in aqueous core (hydrophilic) or within bilayer (hydrophobic). |
| Typical Drug Loading Capacity | 10-30% w/w (can be higher for matrix systems). | 1-10% w/w (limited by core volume/bilayer solubility). |
| Surface Modification | Covalent conjugation or adsorption of ligands, PEGylation via block copolymers. | Post-insertion or co-formulation of functionalized lipids (e.g., PEG, antibodies, peptides). |
| In Vivo Stability | Generally high; degradation controlled by polymer MW & composition. | Variable; can be prone to oxidation, fusion; stabilized by cholesterol/PEG. |
Table 2: Key Characterization Parameters & Techniques
| Characterization Goal | Key Techniques for PNPs | Key Techniques for Liposomes |
|---|---|---|
| Size & Distribution | Dynamic Light Scattering (DLS), SEM, TEM. | DLS, Nanoparticle Tracking Analysis (NTA). |
| Surface Charge | Zeta Potential measurement. | Zeta Potential measurement. |
| Morphology | TEM, SEM, Atomic Force Microscopy (AFM). | Cryo-TEM, AFM. |
| Lamellarity / Structure | Not applicable. | Cryo-TEM, 31P-NMR. |
| Drug Encapsulation | HPLC/UV-Vis after centrifugal ultrafiltration/dialysis. | Mini-column centrifugation, dialysis followed by HPLC/fluorescence assay. |
| In Vitro Release | Dialysis bag method in sink conditions; frequent sampling. | Dialysis or membrane filtration; sampling over time. |
Visualization of Synthesis Workflows
Particle Nanoprecipitation Workflow
Liposome Preparation by Thin-Film Hydration
The Scientist's Toolkit: Essential Research Reagents & Materials
Table 3: Key Reagent Solutions for Nanoparticle Research
| Item | Function/Description | Example Brands/Types |
|---|---|---|
| Biodegradable Polymers | Forms the structural matrix of PNPs; defines degradation kinetics and drug release profile. | PLGA (Lactel, Sigma-Aldrich), PLA, Polycaprolactone (PCL), Chitosan. |
| Phospholipids | Fundamental building blocks of liposome bilayers; determine membrane fluidity and stability. | DPPC, DSPC, POPC, DOPC (Avanti Polar Lipids, CordenPharma). |
| Cholesterol | Incorporated into liposome bilayers to modulate membrane fluidity, permeability, and stability. | High-purity cholesterol (Sigma-Aldrich, Avanti). |
| PEGylated Lipids | Used for "stealth" coating (PEGylation) to reduce opsonization and extend liposome circulation time. | DSPE-PEG2000, DPPE-PEG5000 (Avanti, NOF). |
| Polymeric Stabilizers | Prevent aggregation during PNP formation and in suspension. | Polyvinyl Alcohol (PVA), Poloxamers (Pluronic F68, F127), Polysorbate 80 (Tween 80). |
| Size Exclusion Gels | For purifying liposomes from unencapsulated free drug or small molecules. | Sephadex G-50, Sepharose CL-4B (Cytiva). |
| Extrusion Equipment | For obtaining monodisperse, unilamellar liposomes of defined size. | Polycarbonate membranes, thermobarrel extruder (Avanti, Northern Lipids). |
| Dialysis Membranes | For purification and in vitro drug release studies. | Spectra/Por membranes with appropriate MWCO. |
| Fluorescent Probes | For tracking nanoparticle cellular uptake, biodistribution, and membrane integrity. | DiI, DiO, Rhodamine-PE, Coumarin-6. |
This whitepaper provides an in-depth architectural analysis of liposomes, focusing on their quintessential bilayer membrane structure. This analysis is framed within a broader research thesis investigating the comparative advantages and design principles of polymeric nanoparticles versus liposomes for drug delivery. While polymeric nanoparticles offer robust, tunable matrices from block copolymers, liposomes provide a biocompatible, phospholipid-based mimicry of biological membranes. Understanding the layered, self-assembled structure of liposomes is fundamental to rational design in nanomedicine, enabling researchers to tailor properties like stability, drug loading, and target engagement.
Core Architecture: The Phospholipid Bilayer
The foundational unit of a liposome is the amphiphilic phospholipid molecule, typically featuring a hydrophilic head group (e.g., phosphatidylcholine) and hydrophobic fatty acid tails. In aqueous media, these molecules spontaneously self-assemble into a closed, spherical bilayer, minimizing the energy of the system. This results in a lamellar architecture with distinct layers:
- Inner Aqueous Core: Encapsulates hydrophilic therapeutics.
- Inner Leaflet: The layer of phospholipids facing the core.
- Hydrophobic Bilayer Interior: The region formed by the interdigitated fatty acid tails, which can harbor hydrophobic or amphiphathic drugs.
- Outer Leaflet: The layer of phospholipids facing the external environment.
- Aqueous External Environment.
Quantitative Comparison: Liposomes vs. Polymeric Nanoparticles
The table below summarizes key architectural and performance parameters, contextualizing liposomes within the broader thesis on delivery systems.
Table 1: Comparative Analysis of Liposomal and Polymeric Nanoparticle Architectures
| Parameter | Liposomes (Bilayer Membrane) | Polymeric Nanoparticles (Matrix/Solid Core) | Functional Implication |
|---|---|---|---|
| Core Material | Phospholipids (e.g., DPPC, DSPC), Cholesterol | Biodegradable Polymers (e.g., PLGA, PLA, PCL) | Liposomes are biomimetic; polymers offer mechanical strength. |
| Structural Motif | Lamellar (Bilayer) | Solid Matrix / Core-Shell / Micellar | Determines drug localization (aqueous core vs. bilayer vs. polymer matrix). |
| Typical Size Range | 50 nm - 200 nm (for conventional) | 20 nm - 500 nm | Affects circulation time, biodistribution, and EPR effect. |
| Drug Loading Capacity | Moderate (Depends on solubility & bilayer affinity) | High (Can be engineered for high payload) | Influences dosing and administration frequency. |
| Encapsulation Efficiency | ~15-45% for hydrophilic drugs; higher for lipophilic | Often >70% for many actives | Impacts cost and scalability of manufacturing. |
| Surface Modification | PEG-lipids for stealth; targeting ligands | PEGylation; ligand conjugation to polymer termini | Critical for evading immune clearance and active targeting. |
| In Vitro Release Kinetics | Often biphasic (burst then sustained) | Tunable from days to months via polymer chemistry | Dictates therapeutic profile (sustained vs. immediate release). |
| Key Manufacturing Methods | Thin-film hydration, microfluidics | Nanoprecipitation, emulsification-solvent evaporation | Determines batch homogeneity, scalability, and cost. |
Experimental Protocols for Structural Analysis
Protocol 4.1: Dynamic Light Scattering (DLS) for Size and Zeta Potential
- Objective: Determine liposome hydrodynamic diameter, polydispersity index (PDI), and surface charge (zeta potential).
- Materials: Liposome dispersion, suitable cuvette, DLS/Zeta potential analyzer.
- Method:
- Dilute the liposome sample in filtered (0.22 µm) appropriate buffer (e.g., 1 mM KCl for zeta) to achieve a slight opalescence.
- Load into a clean, dust-free cuvette.
- For size: Measure at a 173° backscatter angle at 25°C. Perform minimum 3 runs, 60 seconds each.
- For zeta potential: Use a dedicated folded capillary cell. Measure electrophoretic mobility and convert to zeta potential via Smoluchowski approximation. Perform >10 measurements.
- Analysis: Report Z-average diameter, PDI, and mean zeta potential ± standard deviation.
Protocol 4.2: Asymmetric Flow Field-Flow Fractionation (AF4) with Multi-Angle Light Scattering (MALS)
- Objective: High-resolution separation and absolute size determination of liposome populations.
- Materials: AF4-MALS system, channel membrane (e.g., regenerated cellulose, 10 kDa MWCO), carrier liquid (e.g., Tris buffer).
- Method:
- Equilibrate the AF4 channel with carrier liquid at a constant tip flow.
- Inject 20-100 µL of liposome sample (~5-10 mg/mL lipid).
- Initiate elution with a cross-flow gradient (e.g., 3.0 to 0.0 mL/min over 30 min) to separate species by hydrodynamic size.
- The eluent passes through in-line UV, MALS, and DRI detectors.
- Analysis: MALS data provides radius of gyration (Rg) for each slice. The Rg/Rh ratio offers insight into vesicle structure and conformation.
Protocol 4.3: Cryo-Electron Microscopy (Cryo-EM) for Morphological Validation
- Objective: Direct visualization of bilayer structure, lamellarity, and morphology.
- Materials: Vitrification system (plunger), holey carbon grids, liquid ethane, cryo-TEM.
- Method:
- Apply 3-5 µL of liposome sample to a glow-discharged grid.
- Blot with filter paper for 2-4 seconds to form a thin film.
- Rapidly plunge into liquid ethane to vitrify the sample.
- Transfer under liquid nitrogen to the cryo-TEM.
- Image at ~-180°C using low-dose procedures (e.g., 120 kV, 30,000-60,000x magnification).
- Analysis: Assess bilayer continuity, vesicle size distribution, and presence of multilamellar vs. unilamellar structures.
Visualization of Liposome Architecture and Characterization Workflow
Title: Liposome Bilayer Structure with Functional Components
Title: Liposome Characterization Experimental Workflow
The Scientist's Toolkit: Key Research Reagent Solutions
Table 2: Essential Materials for Liposome Research and Development
| Reagent/Material | Supplier Examples | Function & Rationale |
|---|---|---|
| High-Purity Phospholipids (DPPC, DSPC, DOPC) | Avanti Polar Lipids, Lipoid GmbH | Primary structural lipids defining bilayer phase transition temperature and stability. |
| Cholesterol | Sigma-Aldrich, Avanti Polar Lipids | Modulates membrane fluidity, packing, and stability; reduces permeability. |
| PEGylated Lipids (DSPE-PEG2000) | NOF America, Avanti Polar Lipids | Creates a hydrophilic steric barrier ("stealth" effect) to reduce opsonization and prolong circulation. |
| Functionalized Lipids (e.g., DSPE-PEG-Biotin, Maleimide) | Avanti Polar Lipids, Nanocs | Enables post-formation conjugation of targeting ligands (antibodies, peptides) to the liposome surface. |
| Lipophilic Tracers (DiI, DiD) | Thermo Fisher Scientific | Fluorescent probes for intercalating into the bilayer, enabling in vitro and in vivo tracking. |
| Size Exclusion Chromatography Columns (Sephadex G-50, Sepharose CL-4B) | Cytiva | For separating untrapped (free) drug or dye from liposomes after preparation (purification). |
| Polycarbonate Membranes (50-200 nm pore size) | Whatman, Sterlitech | Used in extrusion apparatus to calibrate and homogenize liposome size to a narrow distribution. |
| Microfluidic Chips (Staggered Herringbone Mixer) | Dolomite, Precision Nanosystems | Enables precise, reproducible, and scalable continuous-flow manufacturing of nanoliposomes. |
The structural paradigm of nanocarriers is foundational to their function. Within the broader thesis comparing the basic structure and composition of polymeric nanoparticles (PNPs) and liposomes, the matrix core of PNPs represents a distinct architectural philosophy. While liposomes are characterized by an aqueous core sequestered by one or more phospholipid bilayers, emulating cellular membranes, PNPs are defined by a solid or semi-solid polymeric matrix core. This core is a three-dimensional network where the drug is physically entrapped, dissolved, or chemically conjugated, governing release kinetics, stability, and biodistribution. This whitepaper provides a technical dissection of the PNP core, detailing its composition, characterization, and the experimental protocols that reveal its properties.
Core Composition & Material Selection
The polymeric matrix core is primarily fabricated from biodegradable and biocompatible polymers. The choice of polymer dictates core morphology, degradation profile, and drug interaction.
Table 1: Common Polymers for Polymeric Nanoparticle Matrix Cores
| Polymer Class | Specific Polymer(s) | Key Properties | Typical Core Structure |
|---|---|---|---|
| Polyesters | Poly(lactic-co-glycolic acid) (PLGA), Poly(lactic acid) (PLA), Poly(ε-caprolactone) (PCL) | Biodegradable (ester hydrolysis), tunable degradation rate (by Mw & copolymer ratio), FDA-approved for many applications. | Dense, solid matrix. |
| Cationic Polymers | Chitosan, Polyethylenimine (PEI) | Mucoadhesive (chitosan), proton-sponge effect for endosomal escape (PEI), often used for nucleic acid delivery. | Swellable, gel-like matrix. |
| Hydrophilic Polymers | Poly(ethylene glycol) (PEG), Poloxamers | Used as copolymers to create stealth coatings (PEG shell) or thermoresponsive cores (Poloxamers). | Often forms a hydrogel-like core or shell. |
| Poly(alkyl cyanoacrylates) | Poly(butyl cyanoacrylate) (PBCA) | Polymerizes in situ, rapid degradation, used for drug delivery to the brain. | Dense, but rapidly eroding matrix. |
| Dendrimers | PAMAM, PPI | Hyperbranched, monodisperse, multivalent surface for functionalization. | Defined, porous core-shell architecture. |
Table 2: Quantitative Comparison: Polymeric Nanoparticle Core vs. Liposome Core
| Feature | Polymeric Nanoparticle (Matrix Core) | Liposome (Aqueous Core) |
|---|---|---|
| Structural Motif | Solid/semi-solid polymeric network. | Aqueous compartment enclosed by lipid bilayer(s). |
| Typical Size Range | 50-300 nm. | 80-200 nm (Small Unilamellar Vesicles). |
| Drug Loading Capacity | 5-30% w/w (high for hydrophobic drugs). | 1-10% w/w (high for hydrophilic drugs in core; lipophilic in bilayer). |
| Primary Loading Method | Entrapment, adsorption, conjugation. | Passive (aqueous core) or active (transmembrane gradient) loading. |
| Release Kinetic Profile | Biphasic (initial burst + sustained release), controlled by polymer erosion/diffusion. | Monophasic rapid release or sustained via gradient maintenance. |
| In Vitro Stability | High physical stability, slow drug leakage. | Lower stability, prone to fusion, aggregation, and leakage. |
| Key Characterization Tools | DSC (Tg), XRD (crystallinity), SEC (Mw degradation). | DSC (lipid phase transition), DLS (size & lamellarity), FRET (integrity). |
Core Characterization: Experimental Protocols
Protocol: Determining Drug Encapsulation Efficiency (EE) and Loading Capacity (LC)
- Objective: Quantify the amount of drug successfully incorporated into the PNP core.
- Materials: Purified PNP dispersion, appropriate solvent (e.g., acetonitrile for PLGA), HPLC system.
- Procedure:
- Total Drug Measurement: Dissolve 1.0 mL of unpurified PNP formulation in 10 mL of solvent to disrupt the matrix. Analyze drug concentration via validated HPLC-UV method. This is the total drug (TD).
- Free Drug Separation: Purify 2.0 mL of PNP dispersion via size exclusion chromatography (e.g., Sephadex G-25 column) or centrifugal ultrafiltration (MWCO 10 kDa). Collect the purified nanoparticles.
- Entrapped Drug Measurement: Dissolve the purified nanoparticle fraction in 10 mL of solvent. Analyze via HPLC. This is the entrapped drug (ED).
- Calculation:
- Encapsulation Efficiency (%) = (ED / TD) x 100.
- Loading Capacity (%) = (Mass of ED / Total mass of lyophilized nanoparticles) x 100.
- Data Interpretation: High EE indicates efficient core entrapment during formulation. LC defines the payload efficiency of the final product.
Protocol: Differential Scanning Calorimetry (DSC) for Core State Analysis
- Objective: Determine the physical state (crystalline, amorphous) of both the polymer matrix and the encapsulated drug within the core.
- Materials: Lyophilized PNP powder, DSC instrument, sealed aluminum pans.
- Procedure:
- Weigh 3-5 mg of lyophilized PNP sample into a DSC pan and seal it.
- Run a heat-cool-heat cycle (e.g., 25°C to 250°C at 10°C/min under N₂ purge).
- Analyze the thermograms for thermal events: Glass Transition Temperature (Tg) of the polymer, melting endotherms of crystalline drug or polymer.
- Data Interpretation: A disappearance of the drug's melting peak in the PNP thermogram indicates amorphous dispersion within the polymeric matrix, which is desirable for enhancing solubility and release.
Visualizing Core Formation & Drug Release Pathways
Title: Nanoprecipitation and Emulsion-Based Core Formation
Title: Drug Release Mechanisms from Matrix Core
The Scientist's Toolkit: Key Research Reagent Solutions
Table 3: Essential Materials for Polymeric Nanoparticle Core Research
| Reagent/Material | Function & Role in Core Architecture |
|---|---|
| PLGA (50:50, acid-terminated) | The benchmark biodegradable polymer. Ratio (LA:GA) & Mw control core degradation rate and drug release profile. |
| PVA (Polyvinyl Alcohol, 87-89% hydrolyzed) | A common stabilizer/emulsifier. Forms a protective interfacial layer during emulsion, affecting core surface properties and drug release. |
| Dichloromethane (DCM) or Ethyl Acetate | Common organic solvents for dissolving hydrophobic polymers. Their volatility and water-miscibility are critical for nanoprecipitation/emulsion. |
| Dialysis Tubing (MWCO 3.5-14 kDa) | For purifying nanoparticles and removing free polymer, drug, and surfactants, ensuring accurate core characterization. |
| Sephadex G-25 Gel Filtration Columns | For rapid, size-based separation of entrapped drug (in core) from free drug, crucial for measuring encapsulation efficiency. |
| Trehalose or Sucrose | Cryoprotectants for lyophilization. Preserve core nanostructure and prevent aggregation during freeze-drying for storage. |
| Fluorescent Probe (e.g., Nile Red, Coumarin-6) | Hydrophobic dyes incorporated into the core to enable visualization (microscopy) and tracking of nanoparticle uptake in vitro. |
| CCK-8 Assay Kit | Standard cell viability assay to evaluate the biocompatibility of the polymeric core and its degradation products. |
The matrix core of polymeric nanoparticles is a sophisticated, engineered space whose architecture fundamentally diverges from the aqueous core of liposomes. Its composition—a dense, degradable polymeric network—confers superior physical stability and sustained release profiles, particularly for hydrophobic agents. Mastery of core fabrication and characterization protocols, as outlined, is essential for rationally designing PNPs that meet specific therapeutic payload and pharmacokinetic demands. This deep architectural understanding enables direct, informed comparisons with liposomal systems, guiding the selection of the optimal nanocarrier platform for a given biomedical challenge.
Within drug delivery research, the choice of nanocarrier is foundational. This guide contextualizes the use of synthetic and natural polymers for Polymeric Nanoparticles (PNPs) within the broader thesis of comparing their basic structure and composition to liposomes. While liposomes are vesicles formed by phospholipid bilayers, encapsulating hydrophilic and hydrophobic drugs within their aqueous core and lipid membrane respectively, PNPs are solid colloidal particles. Their matrix, composed of biodegradable polymers, offers distinct advantages in stability, controlled release profiles, and tunability of polymer chemistry. This document provides an in-depth technical examination of three quintessential PNP building blocks: the synthetic polyesters PLGA and PLA, and the natural polysaccharide chitosan.
Core Polymer Chemistry & Properties
Poly(lactic-co-glycolic acid) (PLGA): A synthetic, biodegradable copolymer of lactic acid and glycolic acid. The degradation rate, crystallinity, and drug release kinetics are directly tunable by altering the lactide:glycolide ratio (e.g., 50:50, 65:35, 75:25, 85:15).
Poly(lactic acid) (PLA): A synthetic, biodegradable polymer derived from lactic acid. It is more hydrophobic and degrades slower than PLGA due to the absence of glycolic acid monomers, leading to more crystalline structures.
Chitosan: A natural, linear polysaccharide composed of randomly distributed β-(1→4)-linked D-glucosamine and N-acetyl-D-glucosamine. It is derived from chitin deacetylation. Its cationic nature (due to primary amine groups) allows for mucoadhesion and permeability enhancement.
Table 1: Key Physicochemical Properties of Core PNP Polymers
| Polymer | Type | Solubility | Key Functional Groups | Degradation Mechanism | Tg (°C, approx.) |
|---|---|---|---|---|---|
| PLGA | Synthetic Copolymer | Organic solvents (acetone, DCM, ethyl acetate) | Ester linkages | Hydrolysis of ester bonds | 40-55 |
| PLA | Synthetic Homopolymer | Organic solvents (chloroform, DCM) | Ester linkages | Hydrolysis of ester bonds | 45-60 |
| Chitosan | Natural Polysaccharide | Dilute acidic aqueous solutions (pH <6.5) | Primary amine, hydroxyl groups | Enzymatic (e.g., lysozyme) | ~105 (dry) |
Synthesis & Formulation Protocols
PNP formulation methodologies are critical for determining particle size, polydispersity index (PDI), drug loading, and encapsulation efficiency.
Standardized Protocol: Emulsion-Solvent Evaporation (for PLGA/PLA)
Objective: Prepare drug-loaded PLGA nanoparticles.
Research Reagent Solutions & Materials:
- PLGA (50:50, acid-terminated): Core polymer matrix. MW: 7-17 kDa for faster release, 38-54 kDa for sustained release.
- Dichloromethane (DCM): Organic solvent to dissolve polymer and hydrophobic drug.
- Polyvinyl Alcohol (PVA, 1% w/v): Aqueous surfactant solution. Stabilizes the oil-in-water emulsion.
- Model Hydrophobic Drug (e.g., Paclitaxel): Active pharmaceutical ingredient.
- Ultrapure Water: Aqueous phase.
- Probe Sonicator: For primary emulsion formation.
- Magnetic Stirrer: For solvent evaporation and hardening.
Methodology:
- Dissolve 50 mg PLGA and 5 mg of the hydrophobic drug in 5 mL of DCM (organic phase).
- Pour the organic phase into 20 mL of 1% PVA aqueous solution.
- Emulsify the mixture using a probe sonicator (e.g., 70% amplitude, 2 min, pulse cycle 5 sec on/2 sec off) over an ice bath to form a primary oil-in-water (O/W) emulsion.
- Transfer this primary emulsion to 100 mL of 0.1% PVA solution under constant magnetic stirring (500 rpm).
- Stir for 4-6 hours at room temperature to allow complete evaporation of the organic solvent and nanoparticle hardening.
- Centrifuge the suspension at 20,000 rpm for 30 min at 4°C. Wash the pellet with water to remove residual PVA and unencapsulated drug.
- Resuspend the final nanoparticle pellet in an appropriate buffer (e.g., PBS) and lyophilize for storage if needed.
Standardized Protocol: Ionic Gelation (for Chitosan)
Objective: Prepare drug-loaded chitosan nanoparticles via ionic crosslinking.
Research Reagent Solutions & Materials:
- Chitosan (low MW, >75% deacetylated): Cationic polymer backbone.
- Tripolyphosphate (TPP, 0.1% w/v): Ionic crosslinker (anionic).
- Acetic Acid (1% v/v): Solvent for chitosan.
- Model Drug (e.g., Insulin): Can be incorporated into either phase.
Methodology:
- Dissolve 20 mg of chitosan in 10 mL of 1% acetic acid solution under magnetic stirring until clear (pH ~4.5).
- Dissolve the model drug in either the chitosan solution or the TPP solution, depending on its charge/solubility.
- Add the TPP solution dropwise (e.g., at 0.5 mL/min using a syringe pump) into the chitosan solution under constant magnetic stirring (600 rpm).
- Continue stirring for 60 minutes to allow nanoparticle formation via electrostatic interaction.
- Centrifuge at 15,000 rpm for 25 min. Wash and resuspend in buffer.
Table 2: Typical Characterization Data for Model PNPs
| Parameter | PLGA NPs (O/W Evaporation) | Chitosan NPs (Ionic Gelation) |
|---|---|---|
| Mean Particle Size | 150 - 250 nm | 80 - 200 nm |
| Polydispersity Index (PDI) | < 0.2 | 0.1 - 0.3 |
| Zeta Potential | -25 to -40 mV (due to PVA/COOH) | +20 to +60 mV (due to NH₃⁺) |
| Encapsulation Efficiency | 60-90% (hydrophobic drugs) | 20-80% (dependent on drug charge) |
| In vitro Release (PBS, 37°C) | Biphasic: burst (20-40% in 24h), sustained (days-weeks) | Faster, often monophasic (hours-days) |
Comparative Diagram: PNP vs. Liposome Structure & Composition
(Diagram 1: PNP vs Liposome Structure Comparison)
Critical Experimental Workflow: PNP Formulation & Characterization
(Diagram 2: PNP Development Workflow)
The Scientist's Toolkit: Essential Research Reagents
Table 3: Key Reagents for PNP Research
| Reagent/Material | Function/Explanation | Typical Specification |
|---|---|---|
| PLGA (50:50) | Benchmark synthetic copolymer; tunable degradation. | Acid-terminated, MW ~24-38 kDa, Lactide:Glycolide 50:50. |
| Chitosan (Low MW) | Cationic natural polymer; enables mucoadhesion. | Deacetylation degree >75%, Viscosity <200 cps. |
| Polyvinyl Alcohol (PVA) | Surfactant & stabilizer for emulsion methods. | 87-89% hydrolyzed, MW 13-23 kDa, for reproducible coatings. |
| Sodium Tripolyphosphate (TPP) | Ionic crosslinker for chitosan nanoparticles. | ≥98% purity, forms stable polyanionic solution. |
| Dichloromethane (DCM) | Common organic solvent for dissolving PLGA/PLA. | Anhydrous, ≥99.8%, for consistent polymer precipitation. |
| Dialysis Tubing (SnakeSkin) | Purification and in vitro drug release studies. | MWCO 10-20 kDa, for separating free drug from PNPs. |
| Dynamic Light Scattering (DLS) Kit | Measures particle size, PDI, and zeta potential. | Includes disposable folded capillary zeta cells & cuvettes. |
| MTT Reagent (Thiazolyl Blue) | Cell viability assay to assess nanoparticle cytotoxicity. | ≥98% purity, standard 5 mg/mL stock solution in PBS. |
This technical guide examines the core compositional building blocks of liposomal membranes—phospholipids and sterols—and their role in defining the physicochemical and biological properties of liposomes. This discussion is framed within a comparative analysis of basic nanoparticle structures, a central thesis in advanced drug delivery research. The fundamental divergence between polymeric nanoparticles and liposomes lies in their core architecture and assembly logic. Polymeric nanoparticles are typically monolithic or matrix-like structures, formed from the aggregation or polymerization of synthetic or natural polymers (e.g., PLGA, chitosan), where the cargo is entrapped within a solid or gel-like core. In stark contrast, liposomes are vesicular structures, self-assembled from amphiphilic lipids in aqueous environments, creating a fluid, lamellar membrane bilayer that encapsulates an aqueous core. This membrane-based architecture, directly derived from biological membranes, offers distinct advantages in biocompatibility, membrane fusion capabilities, and the ability to host both hydrophilic (in the core) and hydrophobic (within the bilayer) agents. The precise selection and ratio of phospholipids (providing the bilayer matrix) and sterols (primarily cholesterol, modulating membrane fluidity and stability) are therefore critical parameters that dictate liposome performance, setting them apart from the polymer-dominated design space of polymeric nanoparticles.
Core Building Blocks: Structure-Function Analysis
Phospholipids
Phospholipids are amphiphilic molecules consisting of a hydrophilic headgroup and hydrophobic fatty acid tails. They spontaneously form bilayers in water, serving as the primary structural component of liposomes.
Common Classes:
- Phosphatidylcholines (PC): Neutral, most common (e.g., DPPC, POPC, HSPC). Provide main bilayer structure.
- Phosphatidylethanolamines (PE): Often neutral, prone to non-bilayer phases; used for fusogenicity (e.g., DOPE).
- Phosphatidylglycerols (PG): Anionic, introduce negative surface charge to prevent aggregation (e.g., DPPG).
- Phosphatidylserines (PS): Anionic, associated with biological signaling (e.g., apoptosis).
Sterols
Cholesterol is the quintessential sterol in liposome formulation. It is intercalated between phospholipid tails, modulating membrane properties without forming a bilayer by itself.
Key Functions of Cholesterol:
- Condenses the phospholipid packing.
- Modulates Fluidity: Reduces membrane fluidity above the phospholipid's phase transition temperature (Tm) and increases it below Tm.
- Enhances Stability: Decreases permeability to small molecules (e.g., encapsulated drugs) and improves bilayer rigidity in biological fluids.
- Inhibits Phase Transitions, preventing leakiness at phase boundaries.
Table 1: Properties of Common Liposomal Phospholipids
| Phospholipid (Abbrev.) | Phase Transition Temp. (Tm °C) | Net Charge at pH 7.4 | Common Chain Length/Saturation | Key Application Rationale |
|---|---|---|---|---|
| Dipalmitoylphosphatidylcholine (DPPC) | 41 | Neutral | C16:0 (saturated) | High Tm; forms rigid, stable bilayers at 37°C. |
| Distearoylphosphatidylcholine (DSPC) | 55 | Neutral | C18:0 (saturated) | Very high Tm; extremely low permeability. |
| Hydrogenated Soy PC (HSPC) | ~52-55 | Neutral | Mixed, saturated | High-stability, long-circulating liposomes. |
| Palmitoyloleoylphosphatidylcholine (POPC) | -2 | Neutral | C16:0, C18:1 (monounsat.) | Fluid bilayer at physiological temps. |
| Dioleoylphosphatidylcholine (DOPC) | -17 | Neutral | C18:1 (monounsat.) | Highly fluid membrane; model studies. |
| Dioleoylphosphatidylethanolamine (DOPE) | ~ -16 | Neutral | C18:1 (monounsat.) | Fusogenic lipid; used with stabilizers. |
| Dipalmitoylphosphatidylglycerol (DPPG) | 41 | Negative | C16:0 (saturated) | Imparts negative surface charge. |
Table 2: Impact of Cholesterol Incorporation on Membrane Properties
| Cholesterol Mol % | Membrane Fluidity | Permeability to Small Molecules | Mechanical Stability (Rigidity) | Resistance to Serum Proteins | Phase Transition Behavior |
|---|---|---|---|---|---|
| 0% (Pure Phospholipid) | Dictated by Tm & saturation. | High, especially near/at Tm. | Lower | Low | Sharp phase transition at Tm. |
| 20-30% | Moderated. Fluid phase slightly condensed. | Significantly reduced. | Increased. | Moderate. | Phase transition broadened. |
| 33-50% (Optimal Range) | "Liquid-ordered" phase achieved. | Minimized. | Maximized; optimal packing. | High (critical for in vivo stability). | Phase transition essentially abolished. |
| >50% | Risk of crystalline cholesterol precipitation. | May increase due to domain formation. | Can decrease due to defects. | Variable, can decrease. | Complex phase behavior. |
Experimental Protocols for Key Characterization
Protocol 1: Determination of Phase Transition Temperature (Tm) via Differential Scanning Calorimetry (DSC) Objective: To characterize the gel-to-liquid crystalline phase transition of phospholipid bilayers. Methodology:
- Liposome Preparation: Dissolve phospholipid (e.g., DPPC) ± cholesterol in organic solvent. Dry under nitrogen to form a thin film. Hydrate film with buffer (e.g., PBS, HEPES) above the expected Tm (e.g., 60°C) with vigorous vortexing to form multilamellar vesicles (MLVs). Subject to 5-10 freeze-thaw cycles (liquid N₂ / warm water bath) for homogenization.
- DSC Measurement: Load sample (~1-2 mg lipid) and reference (buffer) into the calorimeter. Scan across a temperature range (e.g., 20°C to 60°C for DPPC) at a controlled rate (e.g., 1°C/min). Record heat flow.
- Data Analysis: Identify the peak temperature of the endothermic transition as the Tm. Analyze peak width and enthalpy (area under curve). Note the broadening or disappearance of the peak with cholesterol addition.
Protocol 2: Assessment of Membrane Fluidity/Packing via Fluorescence Polarization Objective: To quantify the microviscosity of the lipid bilayer using a hydrophobic probe. Methodology:
- Probe-Labeled Liposome Preparation: Prepare liposomes as above. Incorporate a fluorescent membrane probe (e.g., DPH: 1,6-diphenyl-1,3,5-hexatriene) at a very low molar ratio (e.g., 1:500 probe:lipid) during the organic solvent dissolution step.
- Measurement: Equilibrate samples at desired temperatures. Use a spectrofluorometer with polarizers. Excite at 360 nm, measure emission intensity at 430 nm parallel (I‖) and perpendicular (I⊥) to the polarized excitation.
- Calculation: Calculate anisotropy (r) = (I‖ - I⊥) / (I‖ + 2I⊥). Higher anisotropy indicates lower fluidity (higher microviscosity). Plot anisotropy vs. temperature or cholesterol content.
Protocol 3: In Vitro Serum Stability / Leakage Assay Objective: To evaluate the stability of liposomes and retention of encapsulated cargo in biologically relevant media. Methodology:
- Dye-Loaded Liposome Preparation: Prepare liposomes (e.g., POPC:Chol 55:45) in buffer containing a high concentration of a self-quenching fluorescent dye (e.g., carboxyfluorescein, CF at 100 mM). Remove unencapsulated dye via gel filtration chromatography (e.g., Sephadex G-50).
- Incubation: Incubate dye-loaded liposomes with fetal bovine serum (FBS) or complete cell culture media (e.g., 1:1 v/v) at 37°C with gentle agitation.
- Measurement: At time points (0, 1, 2, 4, 8, 24 h), dilute samples. Measure fluorescence (ex ~492 nm, em ~517 nm) before (Ft) and after (Ftotal) complete lysis of liposomes with a detergent (e.g., Triton X-100).
- Calculation: Calculate % dye retention = [1 - (Ft / Ftotal)] * 100. Compare formulations with and without cholesterol.
Visualizations
Diagram Title: NP vs Liposome Core Structure Comparison
Diagram Title: Rational Liposome Formulation Workflow
The Scientist's Toolkit: Research Reagent Solutions
Table 3: Essential Materials for Liposome Membrane Research
| Reagent / Material | Key Function / Role | Example Vendor(s) |
|---|---|---|
| Synthetic Phospholipids (e.g., DPPC, POPC, DSPC, DOPE) | High-purity, defined acyl chains provide reproducible bilayer matrix. Essential for structure-function studies. | Avanti Polar Lipids, Merck, CordenPharma |
| Cholesterol (Pharmaceutical Grade) | Modulates membrane fluidity, stability, and permeability. Critical for in vivo applications. | Avanti Polar Lipids, Sigma-Aldrich, NOF America |
| PEGylated Lipids (e.g., DSPE-mPEG2000) | Conjugate to create a hydrophilic polymer (PEG) coat. Confers "stealth" properties by reducing opsonization and extending circulation time. | Avanti Polar Lipids, NOF America, Laysan Bio |
| Fluorescent Membrane Probes (DPH, Laurdan, NBD-PE, Rhodamine-PE) | Report on membrane fluidity, phase, packing, and fusion/rupture events. | Thermo Fisher, Avanti Polar Lipids, Sigma-Aldrich |
| Self-Quenching Dyes (Carboxyfluorescein, Calcein) | Encapsulated at high concentration to assay membrane integrity and leakage kinetics. | Thermo Fisher, Sigma-Aldrich |
| Liposome Extrusion Kit | Equipment (hand-held extruder, membranes) to produce uniform, size-controlled liposomes via membrane filtration. | Avanti Polar Lipids, Cytiva |
| Size Exclusion Chromatography Columns (e.g., Sephadex G-50) | For purifying liposomes from unencapsulated dyes, drugs, or free polymers. | Cytiva, Bio-Rad |
| Differential Scanning Calorimetry (DSC) Instrument | Directly measures the enthalpy and temperature of phospholipid phase transitions. | TA Instruments, Malvern Panalytical |
| Dynamic Light Scattering (DLS) / Zeta Potential Analyzer | Measures liposome hydrodynamic diameter, polydispersity index (PDI), and surface zeta potential. | Malvern Panalytical, Horiba, Beckman Coulter |
Within the context of polymeric nanoparticle (PNP) and liposome research, a fundamental thesis posits that the basic structure and composition are the primary determinants of function. This whitepaper explores how the inherent material properties of these nanocarriers, dictated by their chemical building blocks, govern their physical characteristics, stability, biodistribution, and therapeutic efficacy. The deliberate selection of lipids or polymers is not merely a starting point but the central engineering decision that defines subsequent behavior in biological systems.
Core Composition and Resulting Properties
The chemical identity of the core materials directly defines the nanocarrier's essential character. The table below summarizes key compositional elements and their direct impact on critical properties.
Table 1: Composition-Property Relationships in Nanocarriers
| Compositional Element | Polymeric Nanoparticle (e.g., PLGA, PLA) | Liposome (e.g., Phosphatidylcholine, Cholesterol) |
|---|---|---|
| Core Chemical Bond | Covalent (ester, amide) | Non-covalent (hydrophobic, van der Waals) |
| Formation Driving Force | Solvent displacement, polymerization | Hydrophobic self-assembly in aqueous media |
| Dominant Physical Property | Rigid, high tensile strength, glass transition (Tg) dependent | Fluid to rigid bilayer based on lipid phase transition (Tm) |
| Degradation Mechanism | Hydrolysis / enzymatic cleavage of backbone (controlled) | Phospholipase action, bilayer destabilization (variable) |
| Typical Drug Loading | Entrapment within matrix / covalent conjugation | Encapsulation in aqueous core / insertion into bilayer |
| Surface Modification | Chemical grafting (e.g., PEGylation) requires reactive groups | Post-insertion or pre-formation of functionalized lipids |
Quantitative Performance Dictated by Composition
Performance metrics in drug delivery are quantifiably linked to material choices. The following table compares measurable outcomes.
Table 2: Quantitative Performance Comparison
| Performance Metric | Typical Polymeric Nanoparticle Range | Typical Liposome Range | Key Compositional Driver |
|---|---|---|---|
| Encapsulation Efficiency (%) | 30-70% (hydrophobic drugs) | 50-90% (hydrophilic drugs in core) | Polymer-drug affinity / Lipid bilayer partitioning |
| Particle Size (nm) | 80-250 nm | 70-150 nm (SUV), 100-1000 nm (MLV) | Polymer M.W., synthesis method / Hydration & extrusion parameters |
| Zeta Potential (mV) | -30 to +30 (depends on end group) | -50 to -10 (for anionic PC lipids) | Terminal functional groups / Lipid head group charge |
| Drug Release Half-life (in vitro) | Days to weeks (biphasic) | Hours to days (burst then sustained) | Polymer crystallinity & M.W. / Bilayer fluidity & cholesterol % |
| In Vivo Circulation Half-life | Hours to days (PEGylated) | 10-20 hours (Stealth liposomes) | PEG density & chain length / PEG-lipid concentration |
Experimental Protocols for Characterizing Material-Dictated Behavior
Protocol 1: Determining Phase Transition Temperature (Tm) for Lipids
Objective: To measure the lipid bilayer phase transition, which dictates membrane fluidity and stability.
- Sample Preparation: Prepare a 1 mM liposome suspension in buffer using thin-film hydration and extrusion through a 100 nm membrane.
- Instrumentation: Use a Differential Scanning Calorimeter (DSC).
- Procedure: Load sample and reference (buffer) into calorimeter cells. Scan from 10°C to 60°C at a rate of 1°C/min.
- Analysis: The Tm is identified as the peak of the endotherm on the DSC thermogram. Incorporation of cholesterol (>30%) will broaden and ultimately abolish this peak, indicating modulated membrane rigidity.
Protocol 2: Determining Glass Transition Temperature (Tg) for Polymers
Objective: To characterize polymer backbone rigidity and its implication for drug release kinetics.
- Sample Preparation: Fabricate PNPs via nanoprecipitation. Lyophilize a purified batch to obtain a dry powder.
- Instrumentation: Use a DSC or Dynamic Mechanical Analyzer (DMA).
- Procedure (DSC): Load 5-10 mg of powder. Perform a heat-cool-heat cycle from -20°C to 100°C at 10°C/min. Analyze the second heating cycle.
- Analysis: The Tg appears as a step change in heat capacity. A higher Tg indicates a more rigid polymer matrix, typically leading to slower drug release.
Protocol 3: Critical Micelle/Assembly Concentration (CMC/CAC) Measurement
Objective: To quantify the stability of self-assembled nanostructures upon dilution.
- Sample Preparation: Prepare a series of dilutions from a stock solution of amphiphilic polymer or lipid.
- Probe Method: Use a fluorescent probe like pyrene.
- Procedure: Add a trace amount of pyrene to each dilution. Measure fluorescence emission spectra (excitation at 339 nm).
- Analysis: Plot the intensity ratio of the first (I373) and third (I384) vibrational peaks (I1/I3) vs. log concentration. The inflection point is the CMC/CAC. Lower values indicate greater assembly stability upon intravenous injection dilution.
Diagram 1: Composition-Property-Performance Relationship Flow
Diagram 2: Iterative Research Workflow for Thesis Validation
The Scientist's Toolkit: Key Research Reagent Solutions
Table 3: Essential Materials for Nanoparticle Composition Research
| Reagent / Material | Function & Rationale | Example in Research |
|---|---|---|
| PLGA (Poly(lactic-co-glycolic acid)) | Biodegradable polyester copolymer; backbone hydrolysis rate tunable by LA:GA ratio. Forms solid, drug-encapsulating matrix. | PNP Core: The workhorse polymer for controlled release. 50:50 ratio degrades faster than 75:25. |
| DSPC (1,2-distearoyl-sn-glycero-3-phosphocholine) | Saturated phospholipid with high phase transition temp (~55°C). Provides a rigid, low-permeability bilayer for stable liposomes. | Liposome Bilayer: Key component of long-circulating (e.g., Doxil) liposomes for reduced leakage. |
| mPEG-DSPE (Methoxy PEGylated distearoyl phosphatidylethanolamine) | Amphiphilic polymer-lipid conjugate. PEG corona provides steric stabilization ("stealth" effect) against opsonization. | Surface Modification: Incorporated into PNPs (as micelles) or liposomes to prolong circulation half-life. |
| Cholesterol | Sterol lipid that modulates membrane properties. Incorporation increases bilayer packing, reduces fluidity, and inhibits drug leakage. | Bilayer Modifier: Used at 30-45 mol% in liposomes to enhance in vivo stability. |
| Pyrene | Fluorescent hydrophobic probe. Its emission spectrum is sensitive to local polarity, enabling CMC/CAC determination. | Critical Concentration Assay: Monitors the formation of hydrophobic nano-domains during self-assembly. |
| DSC Calibration Standards (Indium, Tin) | High-purity metals with known, sharp melting points. Used to calibrate temperature and enthalpy scales in DSC for accurate Tm/Tg measurement. | Instrument Calibration: Essential for obtaining reliable, reproducible thermal property data. |
The thesis that structure and composition dictate function is irrefutably demonstrated in the contrasting yet complementary worlds of polymeric nanoparticles and liposomes. From the covalent bonds of a polymer backbone to the self-assembled hydrophobic tails of lipids, each compositional choice propagates through a cascade of physical and chemical properties, ultimately defining biological fate and therapeutic utility. Mastery of these inherent material properties remains the cornerstone of rational nanocarrier design.
From Bench to Bedside: Synthesis Techniques and Therapeutic Applications of Nanocarriers
This technical guide examines key fabrication methods for liposomes, a cornerstone nanocarrier system. Within the broader research comparing the basic structure and composition of polymeric nanoparticles versus liposomes, understanding precise fabrication is critical. Liposomes, with their amphiphilic phospholipid bilayers, mimic biological membranes, offering distinct advantages in biocompatibility and drug encapsulation of hydrophilic and hydrophobic agents. This contrasts with polymeric nanoparticles, which are typically solid matrices formed from synthetic or natural polymers, offering different degradation profiles and mechanical stability. The chosen fabrication method directly dictates critical quality attributes (CQAs) of liposomes—size, polydispersity index (PDI), lamellarity, and encapsulation efficiency—thereby influencing their performance in drug delivery applications relative to their polymeric counterparts.
Thin-Film Hydration (TFH) Method
Detailed Experimental Protocol
Principle: Lipids are dissolved in an organic solvent, which is evaporated to form a thin lipid film. Subsequent hydration with an aqueous buffer leads to the spontaneous formation of multilamellar vesicles (MLVs).
Materials:
- Phospholipids (e.g., DPPC, DSPC, POPC, Cholesterol)
- Organic solvent (chloroform, dichloromethane, or chloroform:methanol mixture)
- Round-bottom flask
- Rotary evaporator connected to a vacuum pump
- Water bath (temperature set above lipid transition temperature, Tc)
- Hydration buffer (e.g., PBS, HEPES, possibly containing the drug for active loading)
Procedure:
- Dissolution: Accurately weigh the lipid components (e.g., 70 mol% phospholipid, 30 mol% cholesterol) and dissolve in organic solvent in a round-bottom flask.
- Film Formation: Attach the flask to a rotary evaporator. Rotate at 40-60 rpm in a water bath above the lipid Tc while gradually applying vacuum. Continue until a smooth, dry lipid film is formed on the inner wall of the flask (typically 30-60 minutes).
- Drying: Place the flask under high vacuum (desiccator) for several hours or overnight to remove trace organic solvent.
- Hydration: Add the pre-warmed (above Tc) aqueous hydration buffer to the flask. The volume determines the final lipid concentration (e.g., 10-20 mM total lipid). Manually swirl or use a rotary evaporator (without vacuum) above Tc for 30-60 minutes to hydrate the film and form a heterogeneous suspension of MLVs.
- Post-processing: The resulting MLV suspension is typically processed further (e.g., by extrusion or sonication) to reduce size and lamellarity.
Extrusion Method
Detailed Experimental Protocol
Principle: A polydisperse liposome suspension (e.g., from TFH) is mechanically passed under pressure through polycarbonate membranes with defined pore sizes, yielding large unilamellar vesicles (LUVs) with a narrow size distribution.
Materials:
- Extruder (hand-held or thermobarrel type, e.g., from Avanti Polar Lipids)
- Polycarbonate membranes (e.g., 100 nm, 200 nm pore size)
- Syringes (typically two)
- Support filters (for membrane)
- Heating block or water bath (for thermosensitive lipids)
Procedure:
- Preparation: Assemble the extruder according to manufacturer instructions. Place a polycarbonate membrane (e.g., 100 nm) between its two halves, supported by appropriate filters. Pre-heat if necessary.
- Loading: Load the MLV suspension (pre-equilibrated above Tc) into one syringe, attach it to one side of the extruder, and place an empty syringe on the opposite side.
- Extrusion: Gently push the suspension through the membrane to the empty syringe. This constitutes one pass. Reverse the syringes and repeat. Typically, a minimum of 11-21 passes are required to achieve a homogeneous, narrow size distribution. The first 1-2 passes may be done through a larger pore size (e.g., 400 nm) to pre-filter the sample.
- Collection: After the final pass, collect the homogeneous LUV suspension from the syringe. The final mean diameter is typically slightly larger than the nominal pore size.
Microfluidics Method
Detailed Experimental Protocol
Principle: Using a microfluidic chip, an aqueous phase and a lipid-containing alcohol phase are mixed in a controlled, rapid manner via hydrodynamic flow focusing or staggered herringbone micromixers. This induces nanoprecipitation, forming liposomes in a single, continuous step.
Materials:
- Microfluidic chip (e.g., planar glass/silicon or PDMS chip with specific mixer design).
- Precision syringe pumps (two or more).
- Phospholipids dissolved in an alcohol (e.g., isopropanol, ethanol).
- Aqueous buffer (e.g., PBS).
- Collection vial.
- Tubing and connectors.
Procedure:
- Solution Preparation: Prepare the lipid stream by dissolving lipids in alcohol (e.g., 10 mM total lipid in ethanol). Prepare the aqueous buffer stream.
- Chip Priming: Flush the microfluidic channels with the respective solvents (alcohol for lipid channels, water for aqueous channels) to remove air bubbles.
- Setup: Load the lipid solution and aqueous buffer into separate syringes. Mount on syringe pumps and connect via tubing to the respective inlets of the microfluidic chip. Place a collection vial at the outlet.
- Flow Rate Optimization: Set the flow rate ratios (FRR). A typical Total Flow Rate (TFR) is 1-3 mL/min with an Aqueous:Organic Flow Rate Ratio (FRR) of 3:1 to 5:1. Higher FRR generally yields smaller liposomes.
- Run & Collection: Start the pumps. The streams meet in the mixing region, causing instantaneous lipid self-assembly into liposomes. The milky suspension is collected continuously from the outlet. The product may be dialyzed or diafiltrated to remove residual alcohol.
Table 1: Comparison of Key Fabrication Methods for Liposomes
| Parameter | Thin-Film Hydration | Extrusion | Microfluidics |
|---|---|---|---|
| Primary Product | Multilamellar Vesicles (MLVs) | Large Unilamellar Vesicles (LUVs) | Monodisperse Unilamellar Vesicles |
| Typical Size Range | 100 nm - 10 µm (pre-extrusion) | 50 nm - 200 nm | 20 nm - 200 nm |
| Polydispersity Index (PDI) | High (>0.3) | Low (<0.1) | Very Low (<0.05) |
| Encapsulation Efficiency (Hydrophilic) | Moderate-Low (passive) | Moderate (passive) | Low-Moderate (passive) |
| Process Scalability | Good for lab-scale batch | Limited by membrane area | Excellent (continuous) |
| Key Advantage | Simple, universal, high drug:lipid ratio possible | Excellent size control, unilamellar | Continuous, tunable, monodisperse |
| Key Limitation | Heterogeneous, requires downstream processing | Batch process, membrane clogging | Dilute suspensions, solvent residue |
Table 2: Impact of Process Parameters on Liposome Characteristics
| Method | Critical Process Parameter | Typical Value/Range | Effect on Liposome CQA |
|---|---|---|---|
| TFH | Hydration temperature | > Lipid Tc | Completeness of film hydration, size of MLVs |
| TFH | Hydration buffer volume & agitation | 1-10 mL, 30-60 min | Lipid concentration, encapsulation efficiency |
| Extrusion | Membrane pore size (nm) | 50, 100, 200 | Final mean particle size (≈1.1-1.2 x pore size) |
| Extrusion | Number of passes | 11-21 | Decreases PDI, increases unilamellarity |
| Extrusion | Pressure/Temperature | ~500 psi, >Tc | Prevents membrane damage, ensures lipid fluidity |
| Microfluidics | Total Flow Rate (TFR, mL/min) | 1-12 | Affects mixing time; influences size & PDI |
| Microfluidics | Flow Rate Ratio (Aq:Org) | 2:1 to 10:1 | Higher ratio → smaller size, lower PDI |
| Microfluidics | Lipid concentration in alcohol | 1-20 mM | Influences final particle size and lamellarity |
The Scientist's Toolkit: Research Reagent Solutions
Table 3: Essential Materials for Liposome Fabrication
| Item | Function/Description | Key Suppliers/Examples |
|---|---|---|
| Phospholipids | Amphiphilic building blocks forming the bilayer. Choices dictate rigidity, charge, and stability. | Avanti Polar Lipids (DOPC, DPPC, DSPC, DMPC), NOF Corporation |
| Cholesterol | Incorporated into bilayers to modulate membrane fluidity, permeability, and stability. | Sigma-Aldrich, Avanti Polar Lipids |
| Polycarbonate Membranes | Porous filters for extrusion defining final liposome size. Available in various pore sizes. | Whatman/Cytiva, Avanti Polar Lipids |
| Liposome Extruder | Device to force liposome suspension through membranes under controlled pressure/temperature. | Avanti Polar Lipids (Mini-Extruder), Northern Lipids |
| Microfluidic Chips | Devices with micron-scale channels for controlled rapid mixing of solvent and aqueous phases. | Dolomite Microfluidics, Micronit, Precision NanoSystems (NanoAssemblr) |
| Rotary Evaporator | For rapid, controlled removal of organic solvent to form thin lipid films in TFH. | Buchi, Heidolph, Yamato |
| Dynamic Light Scattering (DLS) Instrument | Essential for characterizing liposome hydrodynamic diameter, PDI, and zeta potential. | Malvern Panalytical (Zetasizer), Horiba, Beckman Coulter |
Visualized Workflows
Title: Thin-Film Hydration (TFH) Workflow
Title: Liposome Extrusion Process
Title: Microfluidic Liposome Formation
Within the broader research thesis comparing the basic structure and composition of polymeric nanoparticles (PNPs) to liposomes, the choice of fabrication method is paramount. This technical guide focuses on two core techniques: emulsion-solvent evaporation and nanoprecipitation. While liposomes are characterized by their amphiphilic phospholipid bilayers encapsulating an aqueous core, polymeric nanoparticles are defined by their solid, often biodegradable, polymer matrices (e.g., PLGA, PLA, PCL). This structural distinction necessitates fundamentally different fabrication approaches. The methods detailed herein govern critical parameters such as particle size, polydispersity, drug loading efficiency, and release kinetics—all key variables in comparative studies against vesicular liposomal systems for drug delivery applications.
Emulsion-Solvent Evaporation Method
Principle
This method involves dissolving a hydrophobic polymer and the active compound in a water-immiscible organic solvent. This solution is then emulsified in an aqueous phase containing a surfactant to form an oil-in-water (O/W) emulsion. Upon evaporation of the organic solvent, the polymer precipitates, forming solid nanoparticles with the drug entrapped within the matrix.
Detailed Protocol
- Organic Phase Preparation: Dissolve 100-500 mg of polymer (e.g., PLGA 50:50, MW 30,000-60,000 Da) and 5-50 mg of hydrophobic drug in 5-25 mL of volatile organic solvent (e.g., dichloromethane (DCM) or ethyl acetate).
- Aqueous Phase Preparation: Prepare 50-250 mL of an aqueous solution containing a stabilizer (e.g., 0.5-5% w/v polyvinyl alcohol (PVA) or sodium cholate).
- Emulsification: Add the organic phase to the aqueous phase under high-speed homogenization (e.g., 10,000-15,000 rpm for 2-5 minutes using an Ultra-Turrax homogenizer). For smaller particles, this coarse emulsion can be further processed via probe sonication (e.g., 60-80 W for 1-3 minutes on ice) or high-pressure homogenization.
- Solvent Evaporation: Stir the obtained O/W emulsion magnetically at room temperature (or under reduced pressure) for 3-12 hours to allow complete evaporation and diffusion of the organic solvent.
- Purification: Centrifuge the nanoparticle suspension at high speed (e.g., 20,000-25,000 x g for 30-45 minutes), discard the supernatant, and resuspend the pellet in distilled water or a buffer. Repeat 2-3 times to remove residual solvent and free surfactant/drug.
- Lyophilization: The purified nanoparticle suspension can be freeze-dried with a cryoprotectant (e.g., 2-5% w/v trehalose or sucrose) to obtain a stable powder.
Key Process Determinants
- Solvent Choice: Volatility and water-immiscibility (DCM > ethyl acetate).
- Surfactant Type and Concentration: Directly impacts particle size and stability.
- Homogenization Energy/Speed: Inversely correlates with particle size.
Nanoprecipitation (Solvent Displacement) Method
Principle
This technique relies on the interfacial deposition of a polymer following displacement of a semi-polar solvent miscible with water from a lipophilic solution. Upon rapid mixing of the polymer solution with a non-solvent (water), the polymer solubility decreases instantaneously, leading to the precipitation of nanoparticles.
Detailed Protocol
- Organic Phase Preparation: Dissolve 50-200 mg of polymer and 5-30 mg of drug in 5-15 mL of a water-miscible organic solvent (e.g., acetone, acetonitrile, or tetrahydrofuran).
- Aqueous Phase Preparation: Prepare 50-150 mL of water or an aqueous solution containing a stabilizer (e.g., 0.1-1% w/v poloxamer 188 or polysorbate 80). The phase may contain no surfactant.
- Precipitation: Under moderate magnetic stirring (500-800 rpm), inject the organic phase into the aqueous phase using a syringe or peristaltic pump. Rapid, turbulent mixing is critical.
- Solvent Removal: Stir the resulting milky suspension for 1-2 hours to allow for diffusion and evaporation of the organic solvent.
- Purification and Lyophilization: Similar to the emulsion method, centrifuge or dialyze to purify, followed by optional freeze-drying.
Key Process Determinants
- Solvent-to-Non-Solvent Miscibility: Must be high for rapid diffusion.
- Mixing Dynamics: Rate and method of injection govern nucleation and growth.
- Organic-to-Aqueous Phase Volume Ratio: Typically 1:5 to 1:20.
Table 1: Comparative Analysis of PNP Fabrication Methods
| Parameter | Emulsion-Solvent Evaporation | Nanoprecipitation |
|---|---|---|
| Typical Particle Size Range | 80 - 500 nm | 50 - 300 nm |
| Achievable Drug Loading (Theoretical) | Up to 30-40% (High) | Typically 5-20% (Moderate) |
| Entrapment Efficiency (Reported Range) | 40% - 85% | 50% - 95% |
| Preferred Drug Log P | High (Hydrophobic) | Moderate to High |
| Organic Solvent | Water-immiscible (DCM, EA) | Water-miscible (Acetone, THF) |
| Surfactant Requirement | High (Essential for stabilization) | Low to None |
| Key Advantage | High loading capacity, robust for many polymers. | Simple, rapid, narrow size distribution. |
| Key Limitation | High shear stress, residual solvent removal. | Limited to water-miscible solvents and certain polymer-drug combinations. |
Table 2: Common Polymers & Surfactants in PNP Fabrication
| Material Category | Specific Example | Primary Function in PNP Fabrication |
|---|---|---|
| Biodegradable Polymers | Poly(lactic-co-glycolic acid) (PLGA) | Forms the nanoparticle matrix; degrades into lactic/glycolic acid. |
| Biodegradable Polymers | Poly(ε-caprolactone) (PCL) | Slower-degrading, hydrophobic matrix polymer. |
| Stabilizers (Surfactants) | Polyvinyl Alcohol (PVA) | Most common stabilizer in emulsion methods; reduces interfacial tension. |
| Stabilizers (Surfactants) | Poloxamers (e.g., Pluronic F68) | Non-ionic triblock copolymer used in nanoprecipitation; provides steric stabilization. |
| Stabilizers (Surfactants) | Sodium Cholate | Ionic, bile salt surfactant used for small particle formation. |
The Scientist's Toolkit: Key Research Reagent Solutions
Essential Materials for PNP Fabrication Experiments:
- PLGA (50:50, acid-terminated): The benchmark biodegradable copolymer for controlled release PNPs. Function: Provides the structural matrix.
- Polyvinyl Alcohol (PVA, 87-89% hydrolyzed): The industry-standard emulsion stabilizer. Function: Interfacial stabilizer during emulsification; prevents coalescence.
- Dichloromethane (DCM): A volatile, water-immiscible solvent. Function: Dissolves hydrophobic polymers/drugs in the emulsion-solvent evaporation method.
- Acetone (HPLC grade): A water-miscible, semi-polar solvent. Function: Polymer/drug solvent in the nanoprecipitation method.
- Poloxamer 188 (Pluronic F68): A non-ionic, amphiphilic block copolymer. Function: Steric stabilizer in nanoprecipitation, reduces protein adsorption.
- Dialysis Tubing (MWCO 12-14 kDa): For purifying nanoparticles via solvent/impurity diffusion. Function: Removes free drug, surfactants, and organic solvents.
- Trehalose Dihydrate: A non-reducing disaccharide. Function: Cryoprotectant during lyophilization to prevent nanoparticle aggregation upon reconstitution.
- Zetasizer Nano System (or equivalent): Instrument suite utilizing Dynamic Light Scattering (DLS) and Laser Doppler Velocimetry. Function: Measures nanoparticle hydrodynamic diameter, PDI, and zeta potential.
Methodological Workflow Diagrams
Title: Emulsion-Solvent Evaporation Workflow
Title: Nanoprecipitation Method Workflow
Title: PNP vs Liposome Core Characteristics
Within the broader thesis on the basic structure and composition of polymeric nanoparticles (PNPs) versus liposomes, a critical comparative analysis lies in their capacity to encapsulate therapeutic agents. The fundamental architectural dichotomy—a solid polymer matrix versus a phospholipid bilayer enclosing an aqueous core—dictates distinct drug loading paradigms. This guide provides an in-depth technical examination of encapsulation efficiency (EE) strategies for hydrophilic and hydrophobic drugs within these two primary nanocarrier systems, highlighting the interplay between core composition, drug properties, and loading methodology.
Structural Basis for Compartmentalization
Polymeric Nanoparticles (PNPs): Typically composed of biodegradable polymers like PLGA, PLA, or PCL, PNPs form a solid, hydrophobic matrix. Drug incorporation occurs via entrapment within this matrix or surface adsorption. Liposomes: Phospholipid vesicles with one or more concentric bilayers enclosing aqueous compartments. This structure creates distinct environments: a hydrophobic region within the lipid bilayer and hydrophilic compartments in the aqueous core and inter-bilayer spaces.
The inherent compatibility between the drug's solubility profile and the carrier's compartments is the primary determinant of loading efficiency.
Quantitative Comparison of Encapsulation Efficiencies
The following table summarizes typical encapsulation efficiency ranges for different drug-carrier-compartment combinations, based on current literature.
Table 1: Encapsulation Efficiency by Drug Type and Nanocarrier System
| Drug Solubility | Preferred Compartment | Polymeric Nanoparticles (PNPs) | Liposomes | Key Influencing Factors |
|---|---|---|---|---|
| Hydrophobic (e.g., Paclitaxel, Curcumin) | Hydrophobic Matrix / Bilayer | High (70-95%) | Moderate to High (60-85%) | Drug-polymer/lipid affinity, loading method (nanoprecipitation vs. film hydration). |
| Hydrophilic (e.g., Doxorubicin HCl, Cisplatin) | Aqueous Core / Hydrogel Matrix | Low to Moderate (20-50%) | High (up to 90%) with active loading | Core volume, surface charge, use of active loading (pH gradient) for liposomes. |
| Amphiphilic | Interface / Both Compartments | Variable (40-80%) | Variable (50-80%) | Molecular structure, partition coefficient. |
Key Experimental Protocols for Loading and EE Assessment
Protocol: Double Emulsion Solvent Evaporation for Hydrophilic Drugs in PNPs
- Objective: Encapsulate a hydrophilic drug (e.g., protein) within the aqueous core of PLGA nanoparticles.
- Materials: PLGA, PVA (surfactant), dichloromethane (organic solvent), drug in aqueous solution, probe sonicator, magnetic stirrer.
- Method:
- Prepare the primary emulsion (W1/O): Add the aqueous drug solution (W1) to a PLGA solution in DCM (O). Sonicate on ice to form a stable W1/O emulsion.
- Form the double emulsion (W1/O/W2): Pour the primary emulsion into a large volume of aqueous PVA solution (W2). Sonicate again to form the double emulsion.
- Solvent Evaporation: Stir the double emulsion for several hours to evaporate the organic solvent, hardening the polymer and trapping the aqueous droplets.
- Centrifugation & Washing: Collect nanoparticles by ultracentrifugation, wash to remove unencapsulated drug and PVA, and lyophilize.
Protocol: Thin-Film Hydration & Passive Loading for Liposomes
- Objective: Passively load hydrophobic and hydrophilic drugs into liposomal compartments.
- Materials: Phospholipids (e.g., DPPC, cholesterol), chloroform, rotary evaporator, hydration buffer (e.g., PBS), extruder or sonicator.
- Method:
- Film Formation: Dissolve lipids in chloroform in a round-bottom flask. Remove solvent via rotary evaporation to form a thin, dry lipid film.
- Hydration: Hydrate the film with an aqueous buffer (containing the hydrophilic drug for passive loading) above the lipid transition temperature (Tm) with vigorous agitation. This forms multilamellar vesicles (MLVs).
- Size Reduction: Process MLVs through sequential extrusion through polycarbonate membranes (e.g., 100 nm, then 50 nm) to form uniform, small unilamellar vesicles (SUVs).
- Purification: Use size exclusion chromatography or dialysis to separate unencapsulated drug from loaded liposomes.
Protocol: Active (Remote) Loading for Hydrophilic Drugs in Liposomes
- Objective: Achieve high EE for weak base/acid drugs using a transmembrane gradient.
- Materials: Pre-formed "empty" liposomes, ammonium sulfate or citrate buffer (for pH gradient), drug solution.
- Method:
- Prepare liposomes via thin-film hydration using a transmembrane gradient solution (e.g., 250 mM (NH4)2SO4) as the hydration buffer.
- Purify liposomes via dialysis or gel filtration into a neutral, iso-osmotic buffer (e.g., NaCl/Hepes). This creates an ammonium sulfate gradient (acidic inside) or a pH gradient.
- Incubate the purified liposomes with the drug (e.g., doxorubicin) at a temperature above the lipid Tm. The uncharged, permeable drug diffuses across the bilayer, becomes charged in the acidic interior, and is trapped.
- This method can achieve EE > 90%.
Protocol: Standard Encapsulation Efficiency (EE) and Drug Loading (DL) Calculation
- Objective: Quantify the success of a loading procedure.
- Method:
- Separation: Separate nanoparticles from unencapsulated free drug using centrifugation, filtration, or dialysis.
- Lysis/Extraction: For PNPs, dissolve an aliquot in acetonitrile or DMSO. For liposomes, disrupt with Triton X-100 or isopropanol.
- Quantification: Use HPLC or UV-Vis spectroscopy to measure drug concentration in the lysate.
- Calculation:
- EE (%) = (Amount of encapsulated drug / Total initial drug amount) x 100
- DL (%) = (Mass of encapsulated drug / Total mass of nanoparticles) x 100
Visualizing Loading Strategies and Workflows
(Diagram 1: Decision workflow for selecting drug loading strategy based on drug properties and nanocarrier type.)
(Diagram 2: Standard experimental workflow for determining encapsulation efficiency (EE) and drug loading (DL).)
The Scientist's Toolkit: Essential Research Reagents & Materials
Table 2: Key Reagent Solutions for Nanoparticle Drug Loading Studies
| Item | Function & Relevance | Example(s) |
|---|---|---|
| Biodegradable Polymers | Forms the solid, hydrophobic matrix of PNPs. Determines degradation rate and drug release profile. | PLGA (50:50, 75:25), PLA, Poly(ε-caprolactone) (PCL) |
| Phospholipids & Sterols | Building blocks of liposomal bilayers. Chain length and saturation affect membrane fluidity and stability. | DPPC, DSPC, POPC, Cholesterol (for membrane stabilization) |
| Surfactants/Stabilizers | Critical for emulsion stabilization during PNP synthesis and for preventing nanoparticle aggregation. | Polyvinyl Alcohol (PVA), Poloxamers (Pluronic), Polysorbate 80 (Tween 80) |
| Gradient-Forming Agents | Enables active loading in liposomes by establishing transmembrane pH or ion gradients. | Ammonium sulfate, Citrate buffer, Calcium acetate |
| Organic Solvents | Dissolve polymers and lipids for carrier formation. Must be removed to form final nanostructure. | Dichloromethane (DCM), Chloroform, Acetonitrile, Ethyl Acetate |
| Purification Devices | Separate unencapsulated "free" drug from loaded nanoparticles for accurate EE measurement. | Dialysis membranes (MWCO 3.5-14 kDa), Size Exclusion Chromatography columns (Sephadex G-50), Centrifugal Filters |
| Characterization Buffers | Provide a stable, physiological medium for hydrodynamic and zeta potential measurements. | Phosphate Buffered Saline (PBS), HEPES-buffered saline, 10 mM NaCl |
| Lyoprotectants | Preserve nanoparticle integrity and prevent fusion/aggregation during lyophilization for storage. | Trehalose, Sucrose, Mannitol |
This whitepaper details advanced applications of liposomes, framed within a comparative thesis on the basic structure and composition of polymeric nanoparticles versus liposomes. The fundamental distinction lies in material composition: liposomes are closed, spherical vesicles formed by one or more concentric phospholipid bilayers surrounding an aqueous core, while polymeric nanoparticles are typically solid colloidal particles composed of biodegradable polymers like PLGA. This structural difference dictates cargo location (liposomes encapsulate hydrophilic drugs in their core and hydrophobic drugs in the bilayer, while polymeric nanoparticles entrap or adsorb drugs within a polymer matrix), stability, release kinetics, and surface functionalization strategies. The following sections spotlight two pinnacle applications of liposome technology, underpinned by current experimental data and protocols.
Liposomes in Cancer Therapy: The Case of Doxil
Doxil (pegylated liposomal doxorubicin) represents a first-generation, FDA-approved nanomedicine that leverages the Enhanced Permeability and Retention (EPR) effect for passive tumor targeting.
Mechanism of Action and Key Data
The long-circulating, STEALTH characteristics are achieved via surface-grafting with methoxy-polyethylene glycol (PEG). Key quantitative parameters are summarized below.
Table 1: Comparative Properties of Doxil vs. Conventional Doxorubicin
| Property | Doxil (Pegylated Liposome) | Conventional Doxorubicin (Free Drug) |
|---|---|---|
| Plasma Half-life | ~55 hours | ~0.2 hours |
| Volume of Distribution | ~2.8 L | ~254 L |
| Peak Doxorubicin Concentration in Plasma (Cmax) | Significantly higher | Lower |
| Cardiac Uptake | Reduced by >90% | High |
| Palmar-Plantar Erythrodysesthesia (PPE) Incidence | Increased (~20-50%) | Rare |
| Primary Clearance Pathway | Mononuclear Phagocyte System (MPS) | Hepatic metabolism & renal excretion |
Diagram 1: Doxil Structure & EPR-Mediated Delivery Pathway
Key Experimental Protocol: In Vivo Assessment of Liposomal Drug Efficacy in a Xenograft Model
Objective: Evaluate the antitumor efficacy and biodistribution of pegylated liposomal doxorubicin compared to free doxorubicin.
- Tumor Implantation: Subcutaneously inject human cancer cells (e.g., SK-OV-3 for ovarian carcinoma) into the flank of immunodeficient mice (e.g., athymic nude mice).
- Randomization & Dosing: When tumors reach ~100-150 mm³, randomize mice into groups (n=8-10). Administer via tail vein:
- Group 1: Doxil (equivalent to 2-5 mg/kg doxorubicin) in 5% dextrose.
- Group 2: Free doxorubicin (same dose).
- Group 3: Vehicle control (5% dextrose).
- Administer Q7D for 2-3 cycles.
- Monitoring: Measure tumor dimensions (caliper) and body weight 2-3 times weekly. Calculate tumor volume: V = (length × width²)/2.
- Terminal Biodistribution Study: At a defined endpoint (e.g., 48h post-final dose), sacrifice animals. Harvest tumors, heart, liver, spleen, kidneys. Homogenize tissues. Quantify doxorubicin fluorescence (Ex/Em: 470/590 nm) against a standard curve after extraction.
- Statistical Analysis: Compare tumor growth curves (mixed-model ANOVA) and final drug concentrations in tissues (one-way ANOVA with Tukey's post-hoc test).
Liposomes in Vaccine Delivery
Liposomes serve as versatile adjuvants and antigen-delivery vehicles, enhancing humoral and cellular immunity. Modern platforms include virosomes (incorporating viral glycoproteins) and pH-sensitive fusogenic liposomes.
Mechanism and Key Data
Cationic liposomes readily complex with negatively charged mRNA or DNA (forming lipoplexes), while antigens can be encapsulated or surface-conjugated. Key performance data is consolidated below.
Table 2: Liposomal Vaccine Platform Examples & Performance
| Platform | Antigen/Immunogen Type | Key Adjuvant/Component | Immune Response Highlights |
|---|---|---|---|
| Inflexal V (Licensed) | Influenza hemagglutinin & neuraminidase | Virosome (phospholipid + viral proteins) | Robust, long-lasting antibody titers; CD4+ T-cell response. |
| mRNA-LNP (COVID-19 Vaccines) | SARS-CoV-2 Spike protein mRNA | Ionizable cationic lipid, PEG-lipid, cholesterol, phospholipid | High neutralizing antibody titers & strong Th1-biased T-cell response. |
| CAF09 (Experimental) | Subunit protein/peptide | Cationic liposome (DDA/TDB) | Induces potent polyfunctional CD4+ T cells (Th1/Th17). |
Diagram 2: Liposomal Vaccine Immunoactivation Pathway
Key Experimental Protocol: Formulation and In Vivo Evaluation of mRNA-LNP Vaccines
Objective: Formulate and test an mRNA-LNP vaccine for antigen-specific immune responses.
- Microfluidic Formulation:
- Prepare an aqueous phase: mRNA encoding antigen in citrate buffer (pH 4.0).
- Prepare an organic phase: Ionizable cationic lipid (e.g., DLin-MC3-DMA), DSPC, cholesterol, and PEG-lipid in ethanol.
- Use a microfluidic mixer (e.g., NanoAssemblr) to rapidly mix aqueous and organic phases at a defined flow rate ratio (e.g., 3:1 aqueous:organic). This induces spontaneous nanoparticle formation.
- Dialyze the formed LNPs against PBS (pH 7.4) to remove ethanol and raise pH.
- Characterize LNP size (70-100 nm target by DLS), PDI (<0.2), zeta potential (~0-5 mV), and mRNA encapsulation efficiency (RiboGreen assay).
- Mouse Immunization:
- Immunize groups of C57BL/6 mice (n=6-8) intramuscularly with mRNA-LNP (e.g., 1-10 µg mRNA dose) at day 0 and day 21.
- Include controls: saline, empty LNP, and a benchmark vaccine.
- Immune Response Analysis (2 weeks post-boost):
- Humoral Immunity: Collect serum. Measure antigen-specific IgG, IgG1, and IgG2c titers by ELISA.
- Cellular Immunity: Isolate splenocytes. Stimulate with antigen peptides. Perform ELISpot for IFN-γ (Th1/Cytotoxic T cell marker) and IL-4 (Th2 marker). Alternatively, use intracellular cytokine staining for flow cytometry.
The Scientist's Toolkit: Research Reagent Solutions for Liposome Research
Table 3: Essential Materials for Liposome R&D
| Item | Function/Description | Example Vendor/Product |
|---|---|---|
| Phospholipids | Primary structural components forming the bilayer. Choice dictates rigidity, charge, and fusogenicity. | Avanti Polar Lipids: DOPC (neutral), DOTAP (cationic), DOPS (anionic). |
| Cholesterol | Incorporated into bilayers to modulate membrane fluidity and stability, reducing premature leakage. | Sigma-Aldrich, Avanti Polar Lipids. |
| PEGylated Lipids | Conjugated to phospholipids; create a hydrophilic steric barrier to reduce opsonization and extend circulation half-life. | Avanti: DSPE-PEG(2000). NOF: SUNBRIGHT series. |
| Ionizable Cationic Lipids | Critical for mRNA-LNP formulation; positively charged at low pH for complexation, neutral at physiological pH to reduce toxicity. | MedChemExpress: DLin-MC3-DMA, SM-102, ALC-0315. |
| Remote Loading Agents | Ammonium sulfate, citrate gradient) enable active, high-efficiency encapsulation of weak base drugs (e.g., doxorubicin). | MilliporeSigma. |
| Extrusion Equipment | For size homogenization and unilamellar vesicle production via polycarbonate membranes. | Northern Lipids: Extruder, Avanti: Mini-Extruder. |
| Microfluidic Mixers | Enable rapid, reproducible mixing for reproducible nanoparticle synthesis (e.g., LNPs). | Precision NanoSystems: NanoAssemblr. |
| Dynamic Light Scattering (DLS) Instrument | Measures hydrodynamic diameter, polydispersity index (PDI), and zeta potential of nanoparticles. | Malvern Panalytical: Zetasizer. |
| Asymmetric Flow Field-Flow Fractionation (AF4) | High-resolution separation and analysis of nanoparticle populations by size. | Wyatt Technology, Postnova Analytics. |
This analysis is framed within a broader thesis comparing the fundamental structure and composition of polymeric nanoparticles (PNPs) to liposomal carriers. While liposomes are characterized by their amphiphilic phospholipid bilayers enclosing an aqueous core, PNPs are defined by their solid, polymeric matrix (nanospheres) or reservoir systems (nanocapsules). This structural distinction fundamentally dictates their drug loading mechanisms, stability profiles, and interaction with biological systems. PNPs offer superior tunability of polymer composition and degradation kinetics, providing a robust platform for both sustained release and the complex demands of genetic material delivery, which this guide will explore in depth.
Core Technical Principles
Polymer Composition & Synthesis
PNPs for these applications are typically fabricated from biodegradable and biocompatible polymers. Poly(lactic-co-glycolic acid) (PLGA) remains the gold standard for sustained release due to its predictable hydrolysis. For gene delivery, cationic polymers like polyethylenimine (PEI), chitosan, and poly(β-amino esters) (PBAEs) are dominant, facilitating nucleic acid condensation and endosomal escape.
Table 1: Common Polymers for PNPs in Sustained Release vs. Gene Delivery
| Application | Polymer Class | Example Polymers | Key Property | Typical Size Range |
|---|---|---|---|---|
| Sustained Release | Polyesters | PLGA, PLA, PCL | Controlled degradation rate; hydrophobic | 100-250 nm |
| Gene Delivery | Cationic Polymers | PEI, Chitosan, PBAEs | Positive charge density for complexation | 80-200 nm |
| Both | Block Copolymers | PLGA-PEG, PEI-PEG | Stealth properties (PEGylation) | 70-150 nm |
Quantitative Performance Data
Table 2: Comparative Performance Metrics of PNP Formulations
| Parameter | Sustained Release (PLGA) | Gene Delivery (PEI-based) | Measurement Method |
|---|---|---|---|
| Encapsulation Efficiency | 60-85% for small molecules | 90-99% for pDNA/siRNA (complexation) | HPLC, fluorescence assay |
| Release Duration | 5 days to 6 weeks | Typically burst release of complexed cargo | In vitro dialysis in PBS |
| Zeta Potential | -20 to -10 mV (neutral) | +20 to +40 mV (cationic) | Dynamic light scattering |
| Transfection Efficiency | N/A | Often 2-5x higher than liposomes in vitro | Luciferase/GFP reporter assay |
| Cytotoxicity (Cell Viability) | >85% (high) | 60-80% (PEI dose-dependent) | MTT/WST-1 assay |
Detailed Experimental Protocols
Protocol 1: Double Emulsion Solvent Evaporation for Sustained-Release PNPs
This method encapsulates hydrophilic drugs (e.g., proteins) within PLGA nanoparticles.
- Primary Emulsion: Dissolve 50 mg PLGA (50:50 LA:GA) in 2 mL dichloromethane (DIC). Add 0.5 mL of aqueous drug solution (e.g., 10 mg/mL BSA). Sonicate on ice (70% amplitude, 30 s) to form a water-in-oil (w/o) emulsion.
- Secondary Emulsion: Pour the primary emulsion into 10 mL of aqueous polyvinyl alcohol (PVA, 2% w/v). Homogenize at 10,000 rpm for 2 minutes to form a (w/o)/w double emulsion.
- Solvent Evaporation: Stir the double emulsion magnetically for 4 hours to evaporate DIC and solidify nanoparticles.
- Purification: Centrifuge at 21,000 x g for 20 min. Wash pellet 3x with deionized water. Resuspend in buffer for characterization.
- Characterization: Determine size (DLS), morphology (SEM), encapsulation efficiency (microBCA assay of supernatant vs. lysate).
Protocol 2: Ionic Gelation for Chitosan/siRNA NP Formation
A mild, aqueous method for gene delivery PNPs.
- Polymer Solution: Dissolve chitosan (low MW, deacetylated >85%) at 1 mg/mL in sodium acetate buffer (pH 5.5) containing 0.1% (v/v) Tween 80.
- siRNA Solution: Dilute siRNA in nuclease-free water to 0.2 mg/mL.
- Complexation: Under vigorous vortexing, add the siRNA solution dropwise to an equal volume of chitosan solution. Incubate for 1 hour at room temperature to allow polyplex (nanoparticle) formation via electrostatic interaction.
- Crosslinking (Optional): Add 10 μL of 1% (w/v) tripolyphosphate (TPP) solution per mL of nanoparticle suspension. Stir gently for 30 min.
- Purification & Characterization: Purify via ultrafiltration (100 kDa cutoff). Characterize size, zeta potential (DLS), and siRNA binding (gel retardation assay).
Visualization of Key Mechanisms
PNP Gene Delivery Pathway
PNP Sustained Release Mechanism
Synthesis Workflow Comparison
The Scientist's Toolkit: Key Research Reagent Solutions
Table 3: Essential Materials for PNP Research
| Reagent/Material | Supplier Examples | Key Function in PNP Research |
|---|---|---|
| PLGA (50:50) | Lactel (Evonik), Sigma-Aldrich | Benchmark biodegradable polymer for sustained release formulations. |
| Branched Polyethylenimine (bPEI) | Polysciences, Sigma-Aldrich | High cationic charge density for strong nucleic acid complexation; standard for transfection. |
| Chitosan (Low MW, >85% DDA) | NovaMatrix, Sigma-Aldrich | Biocompatible cationic polysaccharide for non-viral gene delivery. |
| Poly(vinyl alcohol) (PVA) | Sigma-Aldrich, MilliporeSigma | Common stabilizer/emulsifier in nanoprecipitation and emulsion methods. |
| Dichloromethane (DCM) | Fisher Scientific, Sigma-Aldrich | Organic solvent for dissolving polymers in emulsion-based synthesis. |
| Tripolyphosphate (TPP) | Sigma-Aldrich | Ionic crosslinker for chitosan nanoparticles. |
| siRNA (e.g., Luciferase/GFP target) | Dharmacon, Sigma-Aldrich | Model nucleic acid for optimization of gene delivery formulations. |
| MTT/XTT Cell Viability Assay Kits | Thermo Fisher, Abcam | Standard colorimetric assay for assessing nanoparticle cytotoxicity. |
| Amicon Ultra Centrifugal Filters | MilliporeSigma | For purification and concentration of nanoparticle suspensions. |
| Dynamic Light Scattering (DLS) System | Malvern Panalytical | Essential instrument for measuring nanoparticle size (hydrodynamic diameter) and zeta potential. |
Within the broader thesis investigating the basic structure and composition of polymeric nanoparticles (PNPs) vs. liposomes, surface functionalization emerges as the critical determinant of in vivo performance and therapeutic efficacy. Both platforms share a common functionalization paradigm centered on two principal strategies: PEGylation for stealth properties and targeting ligand conjugation for active delivery. However, the fundamental differences in their core composition—synthetic polymers versus phospholipid bilayers—dictate distinct chemical approaches, conjugation efficiencies, and final surface architectures. This guide details the technical execution of these strategies for each platform, providing a comparative framework essential for rational nanocarrier design.
Core Chemical Strategies for PEGylation
PEGylation, the covalent attachment or physical incorporation of poly(ethylene glycol) (PEG) chains, creates a hydrophilic corona that reduces opsonization and prolongs systemic circulation.
Polymeric Nanoparticles (PNPs)
PNPs (e.g., PLGA, PLA, PCL) offer surface functional groups (e.g., carboxyl, amine) for post-formation "grafting-to" or permit "grafting-from" approaches during polymerization.
- Grafting-To: Pre-synthesized, end-functionalized PEG (e.g., HOOC-PEG-NHS, NH2-PEG-COOH) is conjugated to surface groups via carbodiimide chemistry (EDC/NHS for -COOH) or glutaraldehyde/ NHS esters for -NH2.
- Grafting-From: Using PEG-containing initiators or monomers (e.g., PEG-PLGA block copolymers) during nanoparticle formulation. This often yields denser, more stable PEG layers.
Liposomes
PEG is predominantly incorporated via "pre-insertion," where lipid-conjugated PEG (PEG-lipid, e.g., DSPE-PEG2000) is mixed with structural lipids before vesicle formation.
- Post-Insertion: PEG-lipids can be incubated with pre-formed liposomes, where the lipid anchor inserts into the bilayer. This is less common for long-circulating designs but useful for specific targeting strategies.
Table 1: Comparative PEGylation Parameters for PNPs vs. Liposomes
| Parameter | Polymeric Nanoparticles (PNPs) | Liposomes |
|---|---|---|
| Primary Method | Grafting-to (post-formation) or Grafting-from (during synthesis) | Pre-insertion (during formulation) |
| Key Reagent | NHS-PEG-COOH, NH2-PEG-NH2, Maleimide-PEG-NHS | DSPE-PEG (e.g., DSPE-PEG2000) |
| Conjugation Chemistry | Carbodiimide (EDC/NHS), Amine-reactive esters | Physical incorporation (hydrophobic anchoring) |
| Typical PEG Density | 5-20 PEG chains per 100 nm² (variable) | 3-10 mol% of total lipid |
| Impact on Size | Moderate increase (~5-15 nm) | Increase of 5-10 nm per PEG2000 layer |
| Critical Consideration | PEG crowding can hinder drug release; batch-to-batch consistency. | PEG-lipid exchange/desorption in vivo ("PEG shedding"). |
Targeting Ligand Conjugation Methodologies
Ligands (antibodies, peptides, aptamers, small molecules) are typically conjugated to the distal end of PEG chains ("PEG-spacer") or directly to the nanoparticle surface.
Experimental Protocol: Maleimide-Thiol Conjugation (for Antibodies/Peptides)
This is a gold-standard, high-efficiency method applicable to both platforms when a maleimide-functionalized surface is present.
A. Surface Activation (for PNPs) or Functional PEG-Lipid Synthesis (for Liposomes)
- PNPs: Synthesize NPs with terminal carboxyl groups. Activate with EDC and Sulfo-NHS in MES buffer (pH 6.0) for 20 min. React with amine-PEG-maleimide (in excess) in PBS (pH 7.4) for 2h. Purify by centrifugation (e.g., 21,000 x g, 20 min) to yield Maleimide-PNP.
- Liposomes: Formulate liposomes containing a functionalized PEG-lipid (e.g., DSPE-PEG2000-Maleimide) at 0.5-1.0 mol% of total lipid. Purify by size-exclusion chromatography (SEC).
B. Ligand Thiolation
- Dissolve antibody (1-2 mg/mL) in PBS (pH 7.4).
- Add 5-10 molar excess of Traut's Reagent (2-Iminothiolane). Incubate for 1h at 4°C.
- Remove excess reagent using a desalting column (e.g., Zeba Spin Column) equilibrated with degassed, EDTA-free buffer.
C. Conjugation Reaction
- Mix thiolated ligand with Maleimide-functionalized nanoparticles (PNPs or liposomes) at a molar ratio optimized for ligand activity (typically 50-100 ligands per particle).
- React under inert atmosphere (N2) for 4-6h at 4°C with gentle agitation.
- Quenching: Add a 100-fold molar excess of L-cysteine relative to maleimide and incubate for 15 min.
- Purification: For PNPs, use centrifugal filtration. For liposomes, use SEC (Sepharose CL-4B column) to remove unreacted ligand. Verify conjugation by SDS-PAGE (shift in ligand MW) or colorimetric assay (e.g., BCA for attached protein).
Alternative Conjugation Chemistries
- Click Chemistry (Cu-free): Strain-promoted alkyne-azide cycloaddition (SPAAC) between DBCO-PEG-lipid/particle and azide-modified ligands. Ideal for sensitive ligands.
- Hydrazone/Carbonyl: Oxidized antibody glycans (periodate treatment) react with hydrazide-functionalized surfaces. Targets Fc region, potentially preserving Fab activity.
Table 2: Ligand Conjugation Efficiency: Representative Data
| Conjugation Method | Platform | Typical Ligand | Conjugation Efficiency (%) | Recommended Characterization |
|---|---|---|---|---|
| Maleimide-Thiol | PNP (PLGA) | Anti-EGFR Fab' | 60-85% | SDS-PAGE, HPLC-SEC |
| Maleimide-Thiol | Liposome (DSPC/Chol) | cRGDfK peptide | 70-90% | Mass Spectrometry, HPLC |
| SPAAC Click | PNP (PCL) | Aptamer | 80-95% | Fluorescence Anisotropy, Gel Shift |
| EDC/NHS (direct) | PNP (PLA-COOH) | Transferrin | 40-70% | Bradford Assay, Zeta Potential Shift |
Diagram: Comparative Functionalization Workflow
(Title: PEGylation and Conjugation Workflow for PNPs and Liposomes)
The Scientist's Toolkit: Essential Research Reagents
Table 3: Key Reagent Solutions for Surface Functionalization
| Reagent/Category | Example Product(s) | Primary Function & Notes |
|---|---|---|
| Functionalized PEGs | Methoxy-PEG-SVA (NHS), Maleimide-PEG-NHS, DSPE-PEG2000-Biotin | Provides stealth layer and chemical handle for subsequent conjugation. Choice of MW (2k-5k Da) and end-group is critical. |
| Coupling Agents | EDC Hydrochloride, Sulfo-NHS | Activates carboxyl groups for amide bond formation with amines. Sulfo-NHS increases water solubility and reaction efficiency. |
| Thiolation Reagents | Traut's Reagent (2-Iminothiolane), SATA (N-Succinimidyl S-Acetylthioacetate) | Introduces sulfhydryl (-SH) groups onto ligands (e.g., antibodies) for maleimide chemistry. SATA allows controlled activation. |
| Click Chemistry Reagents | DBCO-PEG-NHS, Azide-PEG-Lipid, Tetrazine-PEG-Lipid | Enables fast, bioorthogonal conjugation without cytotoxic copper catalysts. Essential for sensitive biomolecules. |
| Purification Tools | Zeba Spin Desalting Columns, Sephadex G-25, Sepharose CL-4B Columns, Amicon Ultra Centrifugal Filters | Removes unreacted cross-linkers, ligands, and byproducts. Critical for conjugate purity and accurate characterization. |
| Characterization Standards | BCA Protein Assay Kit, Ellman's Reagent (DTNB), NHS-Fluorescein | Quantifies ligand density, free thiol groups, and conjugation yield. Fluorescent tags enable FACS or microscopy validation. |
| Lipid Components | DSPE-PEG2000, DSPC, Cholesterol, Maleimide-DPPE | Building blocks for liposome formulation with built-in functionalization capacity. Purity (>99%) is essential for reproducibility. |
Characterization of Functionalized Nanoparticles
Essential post-functionalization analyses include:
- Size & Zeta Potential: Dynamic Light Scattering (DLS) to confirm PEG layer increase (+2-10 nm) and charge masking (zeta potential shift towards neutral).
- Ligand Density: Quantitative methods (e.g., BCA for proteins, HPLC for small molecules, fluorescent calibration curves).
- In Vitro Validation: Cell binding/uptake assays using ligand-receptor positive vs. negative cells, with blockers to confirm specificity.
- Stability & Sterility: Assessment of aggregation (DLS, NTA) over time in physiological buffers and after sterile filtration (0.22 µm).
Successful surface functionalization via PEGylation and ligand conjugation is a multi-step engineering challenge that must be tailored to the intrinsic chemistry of the nanoparticle platform. While liposomes leverage the simplicity of lipid insertion, PNPs offer versatility in covalent grafting strategies. The choice of chemistry, PEG architecture, and ligand attachment point directly influences the pharmacokinetic and pharmacodynamic profiles outlined in the broader thesis. Mastery of these protocols enables the transformation of structurally distinct PNPs and liposomes into targeted, long-circulating nanomedicines.
Overcoming Hurdles: Key Challenges, Stability Issues, and Scalability Solutions
Within the broader thesis comparing the basic structure and composition of polymeric nanoparticles and liposomes, the stability of liposomes emerges as a critical differentiator. While polymeric nanoparticles derive stability from their covalently bonded, solid polymer matrices, liposomes are dynamic, self-assembled structures whose integrity and function are contingent on the chemical and physical stability of their phospholipid bilayers. This guide provides an in-depth technical analysis of three paramount stability challenges for liposomal drug delivery systems: fusion/aggregation, phospholipid oxidation, and ester bond hydrolysis.
Liposome Fusion and Aggregation
Fusion refers to the merging of two or more liposome bilayers into a single entity, while aggregation is the adhesion of vesicles without loss of individual membrane integrity. Both processes increase particle size, potentially leading to rapid clearance and altered biodistribution.
Mechanism: Driven by the hydrophobic effect and van der Waals forces, these processes are accelerated by factors that reduce electrostatic repulsion or hydration forces between vesicles, such as high lipid concentration, freeze-thaw cycles, or the absence of stabilizing charged lipids (e.g., DSPG) or PEG-lipids.
Quantitative Data on Stabilizing Additives:
Table 1: Impact of Formulation Additives on Liposome Fusion/Aggregation
| Additive | Concentration (mol%) | Z-Average Size (nm) After 30 Days at 4°C | Polydispersity Index (PDI) Change | Primary Mechanism |
|---|---|---|---|---|
| None (Pure DPPC) | 0 | 220 → 450 | 0.12 → 0.45 | N/A |
| Cholesterol | 45 | 150 → 155 | 0.08 → 0.10 | Membrane Rigidification |
| DSPG (Anionic) | 10 | 160 → 165 | 0.10 → 0.12 | Electrostatic Repulsion |
| DSPE-PEG2000 | 5 | 155 → 158 | 0.07 → 0.08 | Steric Stabilization |
| Sphingomyelin | 100 | 145 → 148 | 0.06 → 0.08 | High Phase Transition (Tm) |
Experimental Protocol: Monitoring Fusion via FRET
- Objective: Quantify membrane fusion using Förster Resonance Energy Transfer (FRET).
- Materials: Donor lipid (e.g., NBD-DPPE, emission 530 nm), acceptor lipid (e.g., Rhodamine-DPPE, emission 590 nm), purified phospholipids, buffer (e.g., HEPES, pH 7.4), extruder.
- Method:
- Prepare two separate liposome populations: Population A (labeled with 1 mol% each of donor and acceptor) and Population B (unlabeled).
- Mix populations A and B at a 1:1 ratio. The initial high FRET signal is due to proximity of dyes in Population A.
- Upon fusion with unlabeled vesicles, dye dilution occurs, decreasing FRET efficiency.
- Monitor fluorescence emission at 590 nm (acceptor) with excitation at 460 nm (donor) over time and under stress conditions (e.g., temperature cycling).
- Calculate % fusion from the decrease in FRET signal relative to controls (e.g., Triton X-100 lysed for 0% FRET, unmixed Population A for 100% FRET).
Phospholipid Oxidation
Oxidation of unsaturated phospholipid acyl chains (e.g., from soy PC, DOPC) is a radical-mediated chain reaction, producing hydroperoxides and secondary cleavage products like malondialdehyde. This compromises membrane fluidity, increases permeability, and generates cytotoxic byproducts.
Mechanism: Initiated by light, heat, or radical initiators (e.g., Fe²⁺/ascorbate), propagating via peroxyl radicals. Oxidation lowers the main phase transition temperature and increases bilayer permeability.
Quantitative Data on Oxidation Parameters:
Table 2: Oxidation Metrics for Common Phospholipids Under Stress
| Phospholipid | Double Bonds | Lag Time (h) at 40°C in Air | Formation of Conjugated Dienes (μM) at 24h | Relative Permeability Increase (CF leakage) |
|---|---|---|---|---|
| DSPC (18:0) | 0 | >1000 | 0.5 | 1.0x (Baseline) |
| POPC (16:0-18:1) | 1 | 120 | 15.2 | 2.3x |
| DOPC (18:1) | 2 | 48 | 42.1 | 5.7x |
| Soy PC (Mixed) | >2 | 24 | 85.5 | 8.9x |
Experimental Protocol: Quantifying Oxidation via Conjugated Diene Formation
- Objective: Measure primary oxidation products (lipid hydroperoxides) by UV spectroscopy.
- Materials: Phospholipids, oxygenated buffer, UV-transparent cuvettes, water bath/shaking incubator.
- Method:
- Prepare liposomes in oxygen-saturated 10 mM phosphate buffer (pH 7.4).
- Divide sample into aliquots and incubate at a controlled temperature (e.g., 37°C or 50°C) in the dark with constant shaking.
- At defined intervals, withdraw aliquots and immediately dilute in absolute ethanol.
- Record UV absorbance spectra from 200 to 300 nm.
- Calculate the concentration of conjugated dienes using the absorbance at 234 nm and the molar extinction coefficient (ε ≈ 29,500 L mol⁻¹ cm⁻¹).
- Plot concentration versus time to determine the lag phase and propagation rate.
Hydrolysis of Phospholipids
Hydrolysis is the cleavage of ester bonds in phospholipids, catalyzed by H⁺, OH⁻, or specific enzymes. It yields free fatty acids and lysophospholipids, which are destabilizing and lytic. The rate is highly dependent on pH and temperature.
Mechanism: Nucleophilic attack by water on the carbonyl carbon of the sn-1 or sn-2 ester bond. Acid-catalyzed hydrolysis proceeds via protonation of the carbonyl oxygen. Base-catalyzed hydrolysis involves direct attack by hydroxide ion.
Quantitative Data on Hydrolysis Kinetics:
Table 3: Hydrolysis Half-Lives (t1/2) of Phospholipids
| Phospholipid | pH 4.0, 25°C | pH 7.4, 37°C | pH 7.4, 4°C | pH 9.0, 25°C |
|---|---|---|---|---|
| DPPC (Sat.) | ~180 days | ~2.5 years | >10 years | ~60 days |
| POPC (Mono-unsat.) | ~150 days | ~1.8 years | >8 years | ~45 days |
| DMPC (Sat., shorter chain) | ~100 days | ~1.0 year | >5 years | ~30 days |
| With 10 mol% PEG-DSPE | Similar to base lipid | Slightly reduced | Similar | Similar |
Experimental Protocol: Measuring Hydrolysis via Fatty Acid Release
- Objective: Quantify hydrolysis by measuring free fatty acid concentration using a copper-based assay.
- Materials: Liposomes, buffers at target pH, chloroform, methanol, copper reagent (triethanolamine, Cu(NO₃)₂), bathophenanthroline solution, shaking water bath.
- Method:
- Incubate liposome suspensions in buffers of defined pH at controlled temperatures. Include 0.01% NaN³ to prevent microbial growth.
- At time points, extract lipids from aliquots using a Bligh-Dyer (chloroform:methanol) method.
- Evaporate the organic phase under N₂ and redissolve in chloroform.
- Add copper reagent, vortex, and centrifuge. The copper ions complex with free fatty acids in the chloroform layer.
- Add bathophenanthroline reagent to the chloroform layer, which forms a colored complex with copper.
- Measure absorbance at 480 nm. Quantify free fatty acid concentration against a standard curve (e.g., oleic acid).
- Plot fatty acid concentration vs. time to determine hydrolysis rate constants.
The Scientist's Toolkit: Key Research Reagent Solutions
Table 4: Essential Materials for Liposome Stability Research
| Reagent/Material | Supplier Examples | Primary Function in Stability Studies |
|---|---|---|
| High-Purity Phospholipids (e.g., DPPC, DOPC, DSPC) | Avanti Polar Lipids, CordenPharma | Define core bilayer properties; minimize impurity-driven instability. |
| Cholesterol (Pharma Grade) | Sigma-Aldrich, Avanti | Modulates membrane rigidity and permeability; inhibits fusion. |
| PEGylated Lipids (e.g., DSPE-PEG2000) | NOF America, Nanocs | Provides steric stabilization against aggregation and protein adsorption. |
| Fluorescent Probe Lipids (NBD, Rhodamine, DiI) | Thermo Fisher, Avanti | Enables FRET-based fusion assays and membrane trafficking studies. |
| Antioxidants (e.g., α-Tocopherol, BHT) | Sigma-Aldrich | Scavenges free radicals to inhibit lipid oxidation during storage. |
| Chelating Agents (e.g., EDTA, DTPA) | Sigma-Aldrich | Binds transition metal ions (Fe²⁺, Cu²⁺) that catalyze oxidation. |
| Cryoprotectants (e.g., Trehalose, Sucrose) | Pfanstiehl, Sigma-Aldrich | Protects against fusion and leakage during lyophilization. |
| Size Exclusion Chromatography Columns (e.g., Sepharose CL-4B) | Cytiva | Separates intact liposomes from hydrolyzed products/aggregates. |
| Mini-Extruder with Polycarbonate Membranes | Avanti Polar Lipids | Produces monodisperse, defined-size liposomes for reproducible studies. |
| Oxygen Scavenging Systems (e.g., Glucose Oxidase/Catalase) | Sigma-Aldrich | Creates anoxic environments for studying oxidation-sensitive payloads. |
Visual Summaries
Diagram 1: Pathways to Liposome Aggregation and Fusion
Diagram 2: Free Radical Chain Reaction of Lipid Oxidation
Diagram 3: Liposome Stability Study Experimental Workflow
Within the broader comparative research on the basic structure and composition of polymeric nanoparticles (PNPs) versus liposomes, a critical performance differentiator lies in the inherent stability of the polymeric matrix. While liposomes face challenges like phospholipid oxidation and fusion, PNPs are governed by the degradation kinetics of their polymer backbone. This in-depth guide examines the core stability challenge of PNPs: the interplay between polymer degradation kinetics and the resultant, often problematic, initial burst release of encapsulated active pharmaceutical ingredients (APIs). Understanding this relationship is paramount for designing controlled-release PNP formulations that outperform liposomal counterparts in specific therapeutic applications.
Fundamentals: Polymer Degradation Mechanisms
Polymer degradation in PNPs is primarily hydrolytic, occurring via bulk erosion or surface erosion, dictated by polymer crystallinity, hydrophobicity, and molecular weight.
- Bulk Erosion: Water penetrates the entire particle faster than the polymer chains cleave, leading to homogeneous degradation throughout the matrix. This is common in poly(lactic-co-glycolic acid) (PLGA) and often results in a sudden mass loss and potential burst release.
- Surface Erosion: The rate of polymer bond cleavage at the surface exceeds the rate of water penetration into the bulk. The particle shrinks over time while maintaining core integrity, offering more linear release kinetics (e.g., poly(anhydrides)).
Key Degradation Reaction (PLGA Example): The ester bonds in PLGA undergo hydrolytic scission, generating lactic and glycolic acid monomers, which further catalyze degradation (autocatalysis).
Linking Degradation to Burst Release
Burst release is the rapid, initial elution of API from the nanoparticle, often within the first 24 hours. It is linked to degradation kinetics through:
- Surface-Located API: API adsorbed on or near the PNP surface is immediately available.
- Water Influx: Initial hydration creates pores and channels, releasing API via diffusion.
- Initial Degradation Phase: Rapid early-stage cleavage of polymer chains at the surface liberates entrapped API. Faster degradation kinetics exacerbate burst release.
- Polymer Swelling: Hydrophilic polymers swell, increasing mesh size and enabling rapid API diffusion.
Quantitative Relationship: The empirical power law model (Korsmeyer-Peppas) often describes cumulative drug release: M_t / M_∞ = kt^n, where n is the release exponent indicating the mechanism. For PNPs, an initial n ≈ 0.5-1.0 indicates coupled diffusion/degradation leading to burst.
Table 1: Impact of Polymer Properties on Degradation Half-life and Burst Release
| Polymer Type (PVA Stabilized) | Mn (kDa) | LA:GA Ratio (for PLGA) | Degradation Medium (pH) | Degradation T½ (days) | % Burst Release (24h) | Key Reference (Simulated from Current Search) |
|---|---|---|---|---|---|---|
| PLGA | 50 | 50:50 | 7.4 PBS | 20-30 | 30-50% | Kumari et al., 2023 |
| PLGA | 50 | 75:25 | 7.4 PBS | 40-60 | 20-35% | Makadia & Siegel, 2023 Review |
| PLGA | 100 | 50:50 | 7.4 PBS | 35-45 | 25-40% | - |
| PLA | 50 | N/A | 7.4 PBS | 60-120 | 15-30% | - |
| PCL | 80 | N/A | 7.4 PBS | >180 | 10-25% | Dash & Konkimalla, 2012 Review (Updated) |
| Liposome (DPPC/Chol) | N/A | N/A | 7.4 PBS | Stable (>30 days) | <10% (if stable) | Comparative Baseline |
Table 2: Strategies to Modulate Degradation and Minimize Burst Release
| Strategy | Example Implementation | Effect on Degradation Kinetics | Typical Reduction in Burst Release |
|---|---|---|---|
| Polymer Blending | PLGA-PEG or PLGA-PCL blends | Modifies hydration rate & crystallinity; variable. | 20-40% reduction |
| Coating/Shell Layer | PLGA core with a chitosan or lipid shell | Delays water influx; surface erosion dominates initially. | 30-50% reduction |
| Cross-linking | Cross-linked gelatin or chitosan nanoparticles | Slows chain dissolution; increases degradation T½. | 40-60% reduction |
| API-Polymer Conjugation | Drug-polymer prodrug (e.g., Paclitaxel-PLA conjugate) | Release governed by linker hydrolysis, not diffusion. | 50-80% reduction |
| Increased Core Crystallinity | High-crystallinity PLLA vs. amorphous PLGA | Slows water penetration; reduces bulk erosion rate. | 15-30% reduction |
Key Experimental Protocols
Protocol 1: In Vitro Degradation Kinetics Study of PNPs
- Objective: Quantify polymer mass loss and molecular weight change over time.
- Materials: PNP dispersion in PBS (pH 7.4), shake incubator, centrifugal filters (100 kDa MWCO), freeze dryer, GPC/SEC system.
- Procedure:
- Dispense 5 mL of PNP suspension (10 mg/mL) into centrifuge tubes. Incubate at 37°C under gentle agitation.
- At predetermined intervals (e.g., days 1, 3, 7, 14, 28), centrifuge samples (15,000 g, 30 min). Recover the pellet.
- Mass Loss: Wash pellet with water, freeze-dry, and weigh. Mass remaining (%) = (Dry mass_t / Initial dry mass) x 100.
- Molecular Weight: Dissolve the dried pellet in THF or appropriate solvent. Analyze via GPC to determine Mn and Mw reduction.
- pH Monitoring: Measure supernatant pH to track acidic degradation product accumulation (autocatalysis indicator).
Protocol 2: In Vitro Drug Release Profile with Burst Release Quantification
- Objective: Measure cumulative API release, specifically characterizing the initial burst phase.
- Materials: PNP dispersion, dialysis membrane (appropriate MWCO), release medium (PBS with 0.1% w/v Tween 80, pH 7.4), HPLC/UV-Vis.
- Procedure (Dialysis Method):
- Place 2 mL of PNP suspension (equivalent to 1-2 mg API) into a pre-soaked dialysis bag. Seal securely.
- Immerse the bag in 200 mL of pre-warmed (37°C) release medium under sink conditions. Stir at 100 rpm.
- At early time points (1, 2, 4, 8, 12, 24 h), withdraw 1 mL of external medium and replace with fresh pre-warmed medium.
- Analyze sample API concentration via validated HPLC/UV-Vis method.
- Calculate cumulative release, correcting for medium replacement. Burst Release is reported as % released at 24h.
Visualization: Pathways and Workflows
Diagram 1: Degradation-Release Pathway in PNPs
Diagram 2: Degrad & Release Exp Workflow
The Scientist's Toolkit: Key Research Reagent Solutions
Table 3: Essential Materials for PNP Degradation & Release Studies
| Item/Category | Specific Example(s) | Function & Rationale |
|---|---|---|
| Model Polymers | PLGA (50:50, 75:25), PLA, PCL, mPEG-PLGA | Core matrix materials with defined degradation rates and erosion mechanisms. |
| Stabilizer/Surfactant | Polyvinyl Alcohol (PVA), Poloxamer 188, Lecithin | Critical for forming stable emulsions during PNP preparation and preventing aggregation in media. |
| Release Medium | Phosphate Buffered Saline (PBS), with 0.1-0.5% w/v Tween 80 | Maintains physiological pH and ionic strength; surfactant ensures sink conditions for hydrophobic drugs. |
| Separation Tool | Centrifugal Filters (Amicon, 100 kDa MWCO), Dialysis Membranes (10-300 kDa MWCO) | Isolates degraded particles/fragments from soluble polymer chains and released API for analysis. |
| Analytical Standards | Lactic Acid, Glycolic Acid, Monomeric API | Calibration standards for quantifying degradation products and released drug via HPLC. |
| GPC/SEC Standards | Polystyrene or Poly(methyl methacrylate) narrow standards | Essential for calibrating GPC systems to determine polymer molecular weight changes over time. |
| pH Indicator | Micro pH electrode | Tracks acidification of the release medium due to accumulation of acidic degradation products (e.g., from PLGA). |
Within the ongoing research thesis comparing the basic structure and composition of polymeric nanoparticles (PNPs) and liposomes, a central challenge emerges: optimizing the drug payload. Both systems offer distinct advantages for drug delivery, but their fundamental architectures dictate disparate strategies for loading and release. PNPs, with their solid polymeric matrices, excel at encapsulating hydrophobic drugs and can be engineered for sustained release. Liposomes, possessing aqueous cores enclosed by phospholipid bilayers, are ideal for hydrophilic compounds and offer biocompatibility. This guide details advanced, current strategies to maximize drug loading capacity and achieve precise release profiles for both platforms, highlighting their structural determinants.
Strategies to Enhance Loading Capacity
Structural Determinants in Polymeric Nanoparticles
The loading capacity of PNPs is intrinsically linked to polymer-drug compatibility and fabrication methodology.
- Polymer-Drug Affinity: Hydrophobic interactions are primary drivers. Using polymers with functional groups complementary to the drug (e.g., hydrogen bonding, ionic interactions) significantly increases encapsulation efficiency (EE%).
- Nanoprecipitation/Solvent Displacement: The rate of polymer precipitation critically affects drug entrapment. Faster precipitation typically leads to higher EE% for hydrophobic drugs.
- Double Emulsion (W/O/W): Essential for hydrophilic drug loading into PNPs. The stability of the primary water-in-oil emulsion is the key factor determining EE%.
- Prodrug Conjugation: Chemically conjugating the drug to the polymer backbone (creating a polymer-drug conjugate) or to a monomer prior to polymerization achieves near-theoretical 100% loading, as the drug becomes an integral part of the nanostructure.
Structural Determinants in Liposomes
Loading in liposomes is governed by the compartmentalization offered by the bilayer and aqueous core.
- Passive Loading: Relies on drug solubility during formation. Hydrophobic drugs partition into the lipid bilayer, while hydrophilic drugs are encapsulated in the aqueous core. EE% is often limited.
- Active (Remote) Loading: The most powerful strategy for weak acid/base drugs. A transmembrane pH or ion gradient (e.g., ammonium sulfate) is established. The uncharged drug diffuses across the membrane and is trapped inside as a charged, membrane-impermeable species, achieving exceptionally high EE% (>90%).
Table 1: Quantitative Comparison of Loading Strategies
| Strategy | Platform | Typical EE% Range | Key Structural/Compositional Lever |
|---|---|---|---|
| Nanoprecipitation | PNP (hydrophobic drugs) | 60-80% | Polymer solubility, solvent:nonsolvent ratio |
| Double Emulsion | PNP (hydrophilic drugs) | 30-70% | Surfactant type/concentration, homogenization energy |
| Prodrug Conjugation | PNP | ~95-100% | Drug-polymer linker chemistry |
| Passive Loading | Liposome | 10-40% (aqueous core), ~70% (bilayer) | Lipid bilayer rigidity, drug logP |
| Ammonium Sulfate Gradient | Liposome (weak bases) | >90% | Gradient strength, lipid permeability |
Strategies to Control Release Profiles
Tuning Polymeric Nanoparticle Release
Release from PNPs is controlled by diffusion, swelling, and degradation of the polymer matrix.
- Polymer Selection: Fast-degrading polymers (e.g., PLGA low M.W.) yield burst release. Slow-degrading, hydrophobic polymers (e.g., PLA, high M.W. PLGA) provide sustained release.
- Matrix Engineering: Incorporating co-monomers or cross-linkers slows degradation and diffusion. Designing surface-eroding polymers (e.g., polyanhydrides) provides more linear, zero-order release kinetics.
- Stimuli-Responsive Design: Polymers that degrade or change conformation in response to specific triggers (pH, redox, enzymes) enable spatiotemporal control.
Tuning Liposome Release
Release from liposomes is governed by bilayer permeability and stability.
- Lipid Composition: Saturated phospholipids (e.g., DSPC) with high phase transition temperature (Tm) form rigid bilayers for slow release. Unsaturated lipids (e.g., DOPC) create leaky, fluid bilayers.
- Cholesterol Modulation: Cholesterol (up to 50 mol%) condenses the bilayer, reduces permeability, and enhances stability in serum, dramatically slowing release.
- Triggered Release: Using lipids that become permeable upon encountering specific stimuli: pH-sensitive (e.g., DOPE/CHEMS), enzyme-sensitive, or light-sensitive lipids.
Table 2: Quantitative Impact of Composition on Release Kinetics
| System | Composition Modification | Effect on Release Rate (vs. baseline) | Mechanism |
|---|---|---|---|
| PNP | PLGA (50:50, 15kDa) -> PLGA (50:50, 45kDa) | Decrease by ~40% over 7 days | Slower polymer degradation/erosion |
| PNP | PLA -> PLGA (50:50) | Increase (significant burst) | Faster hydrolytic degradation |
| Liposome | DOPC -> DSPC (at 37°C) | Decrease by ~60% over 24h | Increased bilayer rigidity (Tm >37°C) |
| Liposome | Add 40 mol% Cholesterol | Decrease by 70-90% over 24h | Reduced membrane fluidity & permeability |
Detailed Experimental Protocols
Protocol 1: High-Loading PLGA Nanoparticles via Nanoprecipitation
Aim: Encapsulate a hydrophobic drug (e.g., Paclitaxel) with >70% EE.
- Organic Phase: Dissolve 50 mg PLGA (e.g., Resomer RG 503H) and 5 mg drug in 5 mL acetone.
- Aqueous Phase: Prepare 20 mL of 0.5% (w/v) polyvinyl alcohol (PVA, M.W. 30-70 kDa) solution in ultrapure water.
- Formation: Using a syringe pump, inject the organic phase into the stirred aqueous phase (500 rpm) at a rate of 1 mL/min.
- Solvent Removal: Stir for 4 hours at room temperature to evaporate acetone.
- Purification: Centrifuge at 21,000 x g for 30 min, wash pellet, and resuspend in buffer. Filter (0.22 µm) for size analysis (DLS).
- EE% Analysis: Lyse an aliquot of nanoparticles in DMSO. Quantify drug via HPLC against a standard curve. EE% = (Mass of drug in nanoparticles / Initial mass of drug) x 100.
Protocol 2: Active Loading of Doxorubicin into Liposomes via pH Gradient
Aim: Achieve >90% EE of doxorubicin into pre-formed liposomes.
- Liposome Preparation: Hydrate a thin lipid film (e.g., DSPC:Cholesterol:PEG2000-DSPE, 55:40:5 molar ratio) in 300 mM citric acid buffer (pH 4.0). Extrude through 100 nm polycarbonate membranes.
- Establish Gradient: Create a transmembrane pH gradient by exchanging the external buffer to phosphate-buffered saline (PBS, pH 7.4) via dialysis or size-exclusion chromatography.
- Drug Loading: Incubate doxorubicin HCl (at a drug:lipid ratio of 1:10 w/w) with the liposome suspension at 60°C (above Tm of DSPC) for 30-60 minutes.
- Purification: Remove unencapsulated drug via size-exclusion chromatography (Sephadex G-50).
- EE% Analysis: Measure fluorescence of the liposome solution before and after disruption with 1% Triton X-100 (Ex/Em: 470/590 nm). EE% = (Fluorescence after disruption / Total possible fluorescence) x 100.
Visualizing Strategies and Workflows
Diagram Title: PNP Loading Strategy Decision Flow
Diagram Title: Drug Release Control Mechanisms
The Scientist's Toolkit: Essential Research Reagents & Materials
Table 3: Key Reagent Solutions for Payload Optimization Research
| Reagent / Material | Function & Rationale | Typical Vendor/Example |
|---|---|---|
| PLGA (50:50, acid-terminated) | Benchmark biodegradable polymer for PNPs; 50:50 lactide:glycolide ratio offers tunable degradation. | Sigma-Aldrich (Resomer RG 503H), Evonik (Resomer RG 503) |
| DSPC (1,2-distearoyl-sn-glycero-3-phosphocholine) | High-Tm saturated phospholipid for forming stable, low-permeability liposome bilayers. | Avanti Polar Lipids (850365P) |
| Cholesterol (Pharma Grade) | Modulates membrane fluidity and stability; essential component for reducing liposome leakage. | Sigma-Aldrich (C8667), Avanti Polar Lipids (700100P) |
| Ammonium Sulfate Solution (300 mM, pH 4.0) | Creates the transmembrane gradient for active loading of weak base drugs (e.g., doxorubicin) into liposomes. | Prepared in-lab (ultrapure grade salts) |
| mPEG2000-DSPE | PEGylated lipid used to create the steric "stealth" coating on liposomes and some PNPs, prolonging circulation. | Avanti Polar Lipids (880126P) |
| Polyvinyl Alcohol (PVA, M.W. 30-70 kDa) | Common surfactant/stabilizer in PNP nanoprecipitation to control particle size and prevent aggregation. | Sigma-Aldrich (363170) |
| Dichloromethane (DCM) & Acetone (HPLC Grade) | Organic solvents for dissolving polymers and hydrophobic drugs during PNP formulation. | Fisher Scientific, Sigma-Aldrich |
| Mini-Extruder with Polycarbonate Membranes | Equipment for producing uniform, size-defined liposomes and PNPs via membrane extrusion. | Avanti Polar Lipids (610000) |
| Size-Exclusion Chromatography Columns (e.g., Sephadex G-50) | For purifying liposomes and PNPs from unencapsulated drug and free reagents. | Cytiva (GE17-0043-01) |
| Dialysis Membranes (MWCO 12-14 kDa) | Alternative purification method and for studying drug release kinetics in vitro. | Spectrum Labs (132700) |
The transition from benchtop formulations of advanced drug delivery systems, such as polymeric nanoparticles and liposomes, to Good Manufacturing Practice (GMP)-compliant clinical batches represents a critical and challenging phase in therapeutic development. Within the broader thesis on the fundamental structure and composition of polymeric nanoparticles versus liposomes, scalability is the bridge that connects promising in vitro data to viable human trials. This guide details the technical considerations, methodologies, and regulatory frameworks essential for this translation.
Core Structural and Scalability Differences: Polymeric Nanoparticles vs. Liposomes
The inherent structural composition of each system dictates distinct scale-up pathways. Liposomes are phospholipid-based vesicles with aqueous cores, while polymeric nanoparticles are typically solid matrices or nanocapsules composed of biodegradable polymers like PLGA.
Table 1: Structural & Scalability Implications of Nanoparticles vs. Liposomes
| Characteristic | Polymeric Nanoparticles (e.g., PLGA) | Liposomes |
|---|---|---|
| Core Composition | Solid polymer matrix or reservoir system. | Aqueous core surrounded by phospholipid bilayers. |
| Key Scalability Challenge | Reproducible polymer precipitation/solvent removal; residual solvent control. | Precise control of lipid hydration, sizing, and drug loading efficiency. |
| Primary Lab Method | Nanoprecipitation, emulsion-solvent evaporation. | Thin-film hydration, followed by extrusion or sonication. |
| Scalable GMP Unit Operation | High-pressure homogenization, controlled precipitation reactors. | High-shear mixing, tangential flow filtration (TFF), high-pressure extrusion. |
| Critical Quality Attribute (CQA) | Particle size (PDI), drug loading %, polymer degradation rate. | Particle size (PDI), lamellarity, encapsulation efficiency, lipid oxidation. |
| Stability Focus | Hydrolytic degradation kinetics, drug leaching. | Phospholipid oxidation, chemical hydrolysis, vesicle aggregation/fusion. |
A Framework for Scalability: From Lab to GMP
The translation process must be systematic, focusing on identifying and controlling Critical Process Parameters (CPPs) that impact Critical Quality Attributes (CQAs).
Table 2: Scale-Up Considerations for Key Unit Operations
| Unit Operation | Lab Scale (Bench) | Pilot/Clinical Scale (GMP) | Key CPPs to Monitor |
|---|---|---|---|
| Formation/Mixing | Magnetic stirrer, vortex mixer. | High-shear homogenizer, static mixer. | Shear rate, mixing time, energy input. |
| Sizing/Reduction | Probe sonication, manual extrusion. | High-pressure homogenizer, automated extruders. | Pressure, cycle number, temperature. |
| Purification | Differential centrifugation, dialysis. | Tangential Flow Filtration (TFF). | Cross-flow rate, transmembrane pressure, filter pore size. |
| Sterilization | 0.22 µm syringe filtration. | Aseptic processing, sterile filtration skids. | Filter integrity, bioburden, hold times. |
| Lyophilization (if used) | Research-scale freeze-dryer. | GMP lyophilizer with process control. | Freezing rate, primary/secondary drying T/P, cake appearance. |
Experimental Protocols for Process Characterization
Protocol 1: Establishing a Design Space for Nanoprecipitation Scale-Up
Objective: To systematically scale the nanoprecipitation of polymeric nanoparticles from 10 mL to 10 L batch size.
- Lab Scale (10 mL): Prepare polymer and drug solution in organic solvent (e.g., acetone). Using a syringe pump, add this solution dropwise (1 mL/min) to a stirred aqueous phase containing a stabilizer. Record stirring speed (RPM).
- Pilot Scale (1 L): Scale linearly by volume. Utilize a peristaltic pump for controlled addition of the organic phase into a jacketed vessel with mechanical stirring. Monitor temperature.
- Clinical Batch (10 L): Implement a controlled precipitation reactor with high-speed impeller mixing and in-line temperature control. The organic phase is introduced via a dedicated feed line with mass flow control.
- Analysis: At each scale, sample and characterize for particle size (DLS), PDI, and drug loading. Correlate CPPs (addition rate, stir speed, Reynolds number) with CQAs.
Protocol 2: Scaling Active Drug Loading for Liposomes (Remote Loading)
Objective: To achieve consistent >90% encapsulation efficiency for an amphipathic drug during scale-up.
- Lab Scale: Prepare blank liposomes via thin-film hydration and extrusion. Create a transmembrane gradient (e.g., ammonium sulfate). Incubate the drug solution with liposomes at a defined drug-to-lipid ratio, temperature (60°C), and time (30 min).
- Scale-Up: Maintain identical drug-to-lipid ratio, incubation temperature, and time. Replace water bath incubation with a temperature-controlled mixing tank. Ensure mixing is sufficient for homogeneity but not so vigorous as to cause shear-induced degradation.
- Purification: Scale purification using TFF systems. Determine the optimal diavolume for unencapsulated drug removal at both pilot and clinical scales.
- Analysis: Measure encapsulation efficiency via HPLC post-purification at each scale. Monitor liposome integrity and size post-TFF.
Visualization of Key Workflows
Diagram 1: GMP Translation Roadmap
Diagram 2: Liposome GMP Manufacturing Flow
The Scientist's Toolkit: Essential Research Reagent Solutions
Table 3: Key Materials for Nanoparticle & Liposome Translation
| Material/Reagent | Function in R&D | GMP-Grade Consideration for Scale-Up |
|---|---|---|
| PLGA (50:50, varied MW) | Biodegradable polymer matrix for nanoparticle formation. | Must source from GMP-certified vendor with certificates of analysis (CoA) for MW, polydispersity, residual monomers. |
| Hydrogenated Soy Phosphatidylcholine (HSPC) | Primary phospholipid for forming stable, rigid liposome bilayers. | Requires GMP-grade sourcing with strict control of fatty acid composition, oxidation levels. |
| mPEG-DSPE | PEGylated lipid used for creating stealth nanoparticles and liposomes. | GMP-grade with controlled PEG chain length and substitution ratio is critical for reproducible pharmacokinetics. |
| Trehalose | Lyoprotectant for freeze-drying (lyophilization) formulations. | Must be pharmaceutical-grade, endotoxin-controlled, with defined crystalline form. |
| Dichloromethane (DCM) / Acetone | Common organic solvents for polymer dissolution in nanoparticle formation. | Strict residual solvent limits (ICH Q3C). Requires validated solvent removal processes and testing. |
| Ammonium Sulfate | Used to create transmembrane gradients for active drug loading into liposomes. | High-purity, endotoxin-free grade essential. Concentration must be precisely controlled. |
| Tangential Flow Filtration (TFF) Membranes | For diafiltration/concentration of final nano-formulations. | Must be compatible, sterilizable (often single-use assemblies), with validated retention characteristics. |
The efficacy and translational potential of nanomedicines, specifically polymeric nanoparticles (PNPs) and liposomes, are fundamentally governed by their physicochemical stability. A core thesis in comparative nanocarrier research posits that the basic structure and composition of these systems—the hydrophobic polymer matrix of PNPs versus the phospholipid bilayers of liposomes—dictates their inherent vulnerabilities and resilience to sterilization stresses and long-term storage. This guide provides a technical framework for ensuring shelf-life stability, grounded in the structural idiosyncrasies of each system.
Structural Determinants of Stability
The choice between PNPs and liposomes is not merely pharmacological but also logistical. Their inherent stability profiles stem from their composition.
- Polymeric Nanoparticles (PNPs): Composed of synthetic (e.g., PLGA, PLA) or natural polymers, their solid-core matrix provides robust mechanical stability but is susceptible to hydrolytic degradation (especially for polyesters) and Oswald ripening. Sterilization can accelerate polymer chain scission.
- Liposomes: Phospholipid bilayers enclosing an aqueous core. Their stability is a function of membrane integrity, highly sensitive to oxidative degradation of lipids, hydrolysis of ester bonds, and phase transition temperatures. Fusion, aggregation, and leakage are key failure modes.
Sterilization Methodologies: Impact on Structure
Sterilization is a critical, destabilizing unit operation. The chosen method must achieve sterility while minimizing damage to the nanocarrier structure.
Experimental Protocol: Assessing Sterilization Impact
Objective: To evaluate the physicochemical integrity of PNPs and liposomes post-sterilization. Methods:
- Sample Preparation: Prepare identical batches of PNPs (e.g., PLGA) and liposomes (e.g., DSPC/Cholesterol).
- Sterilization Treatments:
- Filtration: Pass through a sterile 0.22 µm polyethersulfone (PES) membrane filter under aseptic conditions.
- Autoclaving: Subject sealed vials to 121°C, 15 psi for 20 minutes.
- Gamma Irradiation: Expose frozen samples to a 25 kGy dose from a ^60^Co source.
- Control: Process aseptically without terminal sterilization.
- Post-Treatment Analysis:
- Size & PDI: Dynamic Light Scattering (DLS) immediately after treatment.
- Chemical Integrity: HPLC for drug content; GC-MS for lipid degradation products (oxidation); GPC for polymer molecular weight change.
- Morphology: TEM or Cryo-EM imaging.
- Leakage (Liposomes): Fluorescence assay using calcein or carboxyfluorescein.
Table 1: Comparative Impact of Sterilization Methods on PNPs vs. Liposomes
| Sterilization Method | Key Parameter | Typical Effect on PNPs | Typical Effect on Liposomes | Structural Rationale |
|---|---|---|---|---|
| Filtration (0.22 µm) | Particle Size (nm) | Minimal change if size << pore size; loss if aggregated. | Potential deformation/fusion under pressure. | Mechanical shear can disrupt labile lipid bilayers; solid polymer core is more resistant. |
| Autoclaving | Size / PDI | Major increase (aggregation, melting). | Drastic increase (fusion, bilayer disruption). | High heat exceeds polymer Tg and lipid Tm, causing structural collapse. |
| Drug Encapsulation | Significant decrease. | Near-total loss. | Enhanced drug diffusion/leakage from destabilized matrix/bilayer. | |
| Gamma Irradiation | Particle Size | Moderate increase possible. | Often stable if antioxidants are present. | Radicals cause polymer cross-linking/scission; lipids undergo peroxidation chain reactions. |
| Chemical Degradation | Reduced Mw (chain scission). | Phospholipid oxidation (increased peroxides). | Radiolysis of water generates hydroxyl radicals that attack polymer chains and unsaturated lipid tails. | |
| Optimal Method | Recommendation | Aseptic processing with terminal 0.22 µm filtration. | Aseptic processing is strongly preferred. | Avoids catastrophic heat/radiation-induced damage to the core nanocarrier structure. |
Storage Stability: Mechanisms and Mitigation
Long-term stability requires controlling the degradation pathways inherent to each structure.
Experimental Protocol: Real-Time & Accelerated Stability Studies
Objective: To determine shelf-life under various storage conditions. Methods:
- Conditions: Aliquot samples and store at:
- 4°C (Refrigeration)
- 25°C / 60% RH (Room Temperature, ICH)
- 40°C / 75% RH (Accelerated, ICH)
- Time Points: Analyze at t=0, 1, 3, 6, 12, 24 months (real-time); 1, 3, 6 months (accelerated).
- Analysis: DLS (size, PDI), Zeta Potential, HPLC (drug content, degradation products), pH, visual inspection (aggregation, precipitation). For liposomes: measure Peroxide Value (PV) and Thiobarbituric Acid Reactive Substances (TBARS).
Table 2: Primary Degradation Pathways and Stabilization Strategies
| Nanocarrier | Primary Degradation Pathway | Critical Quality Attribute (CQA) Affected | Stabilizing Excipient/Strategy | Mechanism of Action |
|---|---|---|---|---|
| PNPs | Hydrolytic Degradation | Molecular Weight, Drug Release Kinetics | Use of end-group capped polymers (e.g., PLGA RG); Lyophilization. | Slows water ingress into polymer matrix, reducing chain scission rate. |
| Aggregation/Oswald Ripening | Particle Size, PDI | Lyoprotectants (Sucrose, Trehalose) during freeze-drying. | Forms amorphous glass, immobilizes particles, prevents fusion. | |
| Liposomes | Lipid Oxidation | Peroxide Value, Membrane Integrity | Antioxidants (α-Tocopherol, BHT); Use of saturated lipids (DSPC). | Scavenges free radicals, terminates peroxidation chain reaction. |
| Hydrolysis | Lysolipid formation, pH decrease | Buffer to neutral pH (e.g., HEPES, pH 7.4); Avoid extreme pH. | Minimizes acid-catalyzed hydrolysis of phospholipid ester bonds. | |
| Fusion & Aggregation | Particle Size, Encapsulation Efficiency | Incorporate cholesterol (>30 mol%); Store above lipid Tm. | Modulates membrane fluidity and mechanical strength, preventing fusion. |
The Scientist's Toolkit: Key Reagents & Materials
Table 3: Essential Research Reagent Solutions for Stability Studies
| Item | Function | Example Use-Case |
|---|---|---|
| Lyoprotectants | Protect nanoparticle structure during freeze-drying by forming a stable glassy matrix. | 5% (w/v) Trehalose or Sucrose in the formulation prior to lyophilization of PNPs. |
| Cryoprotectants | Prevent ice crystal damage and particle aggregation during freezing (pre-lyophilization). | Glycerol or DMSO for liposome storage at -80°C. |
| Antioxidants | Inhibit oxidative degradation of lipids (liposomes) or sensitive drug payloads. | 0.1% (w/v) α-Tocopherol incorporated into the lipid bilayer during liposome prep. |
| Sterile Filtration Membranes | For terminal sterilization of thermally/radiolytically sensitive nanocarriers. | 0.22 µm PES syringe filters for aseptic filtration of small (<200 nm) PNPs. |
| Stability Study Buffers | Maintain pH to prevent acid/base-catalyzed degradation (hydrolysis of PLGA/phospholipids). | 10 mM HEPES buffer, pH 7.4, for long-term liposome storage. |
| Inert Atmosphere | Remove oxygen to prevent oxidation during storage. | Argon or Nitrogen headspace flushing prior to vial crimping for lipid-based systems. |
| Stabilizing Lipids | Enhance liposome bilayer rigidity and reduce permeability/fusion. | Incorporating 40 mol% Cholesterol into DSPC liposomes. |
Decision and Analysis Workflows
Diagram Title: Sterilization Method Decision Workflow
Diagram Title: Nanocarrier-Specific Degradation Pathways
Within the fundamental research comparing the structure and composition of polymeric nanoparticles (PNPs) and liposomes, the strategic selection of raw materials and the inherent complexity of production are pivotal factors. This analysis provides a technical framework for evaluating these considerations, directly impacting research reproducibility, scalability, and therapeutic application viability.
Raw Material Sourcing: Comparative Analysis
Sourcing involves evaluating cost, purity, reproducibility, and supply chain stability for core components.
Table 1: Raw Material Sourcing Profile for Liposomes vs. Polymeric Nanoparticles
| Material Category | Liposomes (Example: Doxil-like) | Polymeric Nanoparticles (Example: PLGA-based) | Key Sourcing Considerations |
|---|---|---|---|
| Primary Structural | Hydrogenated soy phosphatidylcholine (HSPC), Cholesterol, PEG-DSPE | Poly(lactic-co-glycolic acid) (PLGA), PEG-PLGA | Lipid purity (>99%) vs. polymer Mw/end-cap variability, polydispersity index (PDI). |
| Cost (per gram, approx.) | HSPC: $200-500; Cholesterol: $50-100; PEG-DSPE: $10,000-15,000 | PLGA (50:50): $100-300; PEG-PLGA: $500-2,000 | Bulk synthesis reduces polymer cost; functionalized lipids remain premium. |
| Supplier Variability | High (natural sourcing affects lipid chain consistency). | Moderate (synthetic control offers better batch-batch reproducibility). | Need for Certificate of Analysis (CoA) detailing fatty acid chains, oxidation levels. |
| Critical Quality Attribute (CQA) | Phase transition temperature (Tm), peroxide value, critical micelle concentration. | Molecular weight, lactide:glycolide ratio, intrinsic viscosity. | Directly impacts nanoparticle size, drug release kinetics, and in vivo stability. |
Production Complexity: Methodologies and Protocols
Production complexity encompasses the number of steps, required instrumentation, process control, and reproducibility.
Detailed Experimental Protocol: Thin-Film Hydration for Liposomes
Objective: To prepare PEGylated, drug-loaded liposomes. Materials: HSPC, Cholesterol, PEG-DSPE, Chloroform, Drug (e.g., Doxorubicin citrate), Citrate buffer (pH 4.0), Phosphate Buffered Saline (PBS, pH 7.4). Procedure:
- Lipid Film Formation: Dissolve lipids (molar ratio 55:40:5 HSPC:Chol:PEG-DSPE) in chloroform in a round-bottom flask. Evaporate solvent using rotary evaporation (40°C, 60 min) to form a thin lipid film.
- Hydration: Hydrate the dried film with pre-warmed (60°C) citrate buffer (300 mM, pH 4.0) to a final lipid concentration of 10-20 mM. Vortex and agitate at 60°C for 1 hour to form multilamellar vesicles (MLVs).
- Size Reduction: Subject the MLV suspension to 5-10 freeze-thaw cycles (liquid N₂/60°C water bath). Then extrude through polycarbonate membranes (400 nm, then 100 nm) using a thermobarrel extruder (60°C) to form small unilamellar vesicles (SUVs).
- Remote Drug Loading: Incubate extruded liposomes with doxorubicin (drug:lipid ratio 1:10 w/w) at 60°C for 1 hour. The pH gradient drives drug encapsulation.
- Purification: Remove unencapsulated drug via size-exclusion chromatography (Sephadex G-50) or tangential flow filtration, equilibrating with PBS pH 7.4.
Detailed Experimental Protocol: Nanoprecipitation for Polymeric Nanoparticles
Objective: To prepare drug-loaded PLGA-PEG nanoparticles. Materials: PLGA (50:50, 10kDa), PEG-PLGA (5kDa), Acetone, Drug (e.g., Paclitaxel), Polyvinyl alcohol (PVA, 1% w/v), Deionized Water. Procedure:
- Organic Phase Preparation: Dissolve PLGA, PEG-PLGA (9:1 ratio), and drug (10% w/w of polymer) in acetone (5 mL total) under magnetic stirring.
- Aqueous Phase Preparation: Place 20 mL of 1% PVA solution in a beaker under vigorous magnetic stirring (600 rpm).
- Nanoprecipitation: Using a syringe pump, inject the organic phase into the aqueous phase at a controlled rate (1 mL/min). Instantaneous diffusion of acetone into water causes polymer precipitation into nanoparticles with encapsulated drug.
- Solvent Removal: Stir the suspension openly for 4 hours to evaporate acetone, or use rotary evaporation under reduced pressure.
- Purification: Centrifuge the suspension at 20,000 x g for 30 minutes, wash pellet with DI water, and resuspend. Alternatively, use tangential flow filtration for larger volumes.
Table 2: Production Complexity Comparison
| Complexity Factor | Liposomes (Thin-Film Hydration) | Polymeric NPs (Nanoprecipitation) |
|---|---|---|
| Key Steps | Film formation, hydration, extrusion, active loading, purification. | Polymer dissolution, nanoprecipitation, solvent removal, purification. |
| Critical Equipment | Rotary evaporator, thermobarrel extruder, size-exclusion chromatography. | Syringe pump, magnetic stirrer, centrifuge or TFF system. |
| Process Control | Temperature critical (must exceed Tm of lipids). Extrusion pressure affects size. | Injection rate, stirring speed, and surfactant concentration are key. |
| Scalability Path | Tangential Flow Filtration (TFF) for hydration/purification; high-pressure homogenization. | Continuous flow microreactors; scaled-up TFF for purification. |
| Typical Encapsulation Efficiency | High for amphiphilic drugs (e.g., Doxorubicin, >90%). | Variable, often moderate for hydrophobic drugs (e.g., Paclitaxel, 60-80%). |
| Batch-to-Batch Variability | Moderate to High (due to natural lipid sources and film homogeneity). | Low to Moderate (due to synthetic polymer consistency). |
The Scientist's Toolkit: Research Reagent Solutions
Table 3: Essential Materials for Nanoparticle Formulation Research
| Item | Function in Research | Key Consideration |
|---|---|---|
| HSPC (Hydrogenated Soy PC) | Provides main bilayer structure; high phase transition temp ensures in vivo stability. | Source from reputable vendors (e.g., Avanti) with detailed CoA; monitor peroxide value. |
| PLGA (50:50 Lactide:Glycolide) | Biodegradable polyester backbone for PNPs; erosion controls drug release. | Select specific molecular weight (e.g., 10-30 kDa) and end-cap (acid or ester) for desired release profile. |
| PEG-DSPE / PEG-PLGA | Provides steric stabilization ("stealth" effect) to evade immune clearance. | PEG chain length (1k-5k Da) is crucial; affects circulation half-life and potential immunogenicity. |
| Thermobarrel Extruder | Essential for producing monodisperse, unilamellar liposomes via membrane extrusion. | Pore size of membranes (50-400 nm) determines final particle size; temperature control is vital. |
| Syringe Pump | Enables controlled, reproducible injection for nanoprecipitation and microfluidic methods. | Precise flow rate (µL/min to mL/min) determines mixing dynamics and final nanoparticle size. |
| Tangential Flow Filtration (TFF) | System for concentrating and purifying nanoparticle suspensions, exchanging buffers. | Molecular weight cutoff (MWCO) of membrane must be appropriate to retain nanoparticles. |
| Dynamic Light Scattering (DLS) | Instrument for measuring hydrodynamic diameter, PDI, and zeta potential. | Sample must be free of dust/aggregates; measurement in relevant biological buffer is ideal. |
Visualizing Key Pathways and Workflows
Diagram Title: Liposome Production via Thin-Film Hydration
Diagram Title: Polymeric Nanoparticle Production via Nanoprecipitation
Diagram Title: Platform Selection Decision Tree
The cost-benefit analysis between liposomes and polymeric nanoparticles reveals a fundamental trade-off. Liposomes offer superior encapsulation for certain drug classes but introduce complexity and cost in raw material sourcing and active loading. Polymeric nanoparticles, particularly from synthetic polymers like PLGA, offer more predictable sourcing and simpler, scalable production (e.g., nanoprecipitation), but may present challenges with drug loading efficiency. The choice within structural research must be guided by the therapeutic agent's physicochemical properties, the required Critical Quality Attributes (CQAs), and the long-term translational goals of the drug development pathway.
Head-to-Head Evaluation: Stability, Efficacy, Safety, and Regulatory Landscape
Within the ongoing research thesis examining the basic structure and composition of polymeric nanoparticles versus liposomes, a critical evaluation of physical stability is paramount. This in-depth technical guide provides a comparative analysis of thermal, mechanical, and long-term shelf-life stability, which are decisive factors for selecting a nanocarrier platform in pharmaceutical development. Structural integrity under stress directly influences drug encapsulation efficiency, biodistribution, safety, and efficacy.
Core Structural Determinants of Stability
Polymeric Nanoparticles: Typically composed of biodegradable polymers like PLGA, PLA, or chitosan. Their stability is governed by polymer crystallinity, molecular weight, glass transition temperature (Tg), and cross-linking density. The solid matrix provides robust mechanical strength but can be susceptible to polymer degradation (hydrolysis) over time.
Liposomes: Phospholipid bilayers enclosing an aqueous core. Stability is dictated by lipid composition (saturated vs. unsaturated chains), phase transition temperature (Tm), cholesterol content (modulates membrane fluidity and permeability), and surface charge. Their fluidic structure confers different stress-response mechanisms.
Quantitative Stability Comparison
The following tables summarize key stability parameters based on current literature.
Table 1: Thermal Stability Comparison
| Parameter | Polymeric Nanoparticles (e.g., PLGA) | Liposomes (e.g., DPPC/Chol) |
|---|---|---|
| Key Transition Temp. | Glass Transition (Tg): ~45-55°C for PLGA | Phase Transition (Tm): ~41°C for DPPC |
| Effect of Heating | Above Tg: Softening, potential deformation & drug burst release. | Above Tm: Transition from gel to liquid crystalline phase; increased permeability. |
| Stabilizing Additives | Plasticizers (e.g., PEG) to modify Tg; cryoprotectants for freeze-drying. | Cholesterol (↑ Tm & broadens transition); Saturated phospholipids (↑ Tm). |
| Typical Stress Test | Differential Scanning Calorimetry (DSC) to measure Tg. | DSC to measure Tm and enthalpy. Turbidity changes. |
Table 2: Mechanical & Shear Stress Stability
| Parameter | Polymeric Nanoparticles | Liposomes |
|---|---|---|
| Structural Response | Resistant to deformation due to solid matrix. Prone to fracture under extreme shear. | Vesicle deformation, potential fusion or rupture under shear. |
| Aggregation Propensity | Lower under shear, but can occur if surface stabilizer is displaced. | Higher, especially for neutral or electrostatically screened vesicles. |
| Standard Test | High-pressure homogenization; extrusion through defined pores; dynamic light scattering (DLS) pre/post stress. | Sonication; extrusion; capillary shear flow with DLS/SLS monitoring. |
Table 3: Long-Term Shelf-Life Stability (Liquid Formulation)
| Parameter | Polymeric Nanoparticles | Liposomes |
|---|---|---|
| Major Degradation Pathways | Polymer hydrolysis (size ↑, molecular weight ↓), drug leaching. | Lipid oxidation, hydrolysis (lyso-lipid formation), drug leakage, aggregation/fusion. |
| Size Change Indicator | Gradual increase due to swelling/aggregation. | Often a bimodal distribution appears due to fusion. |
| Optimal Storage | Often lyophilized with cryo/lyoprotectants (sucrose, trehalose). | 4°C, inert atmosphere (N2), often lyophilized with trehalose. |
| ICH Accelerated Testing | 40°C/75% RH for 1-6 months; monitoring size, PDI, drug content. | Same conditions, plus monitoring of peroxide value & encapsulation efficiency. |
Experimental Protocols for Stability Assessment
Protocol 1: Accelerated Thermal Stability Study
- Sample Preparation: Prepare identical batches of lyophilized or aqueous suspensions of polymeric NPs and liposomes. Aliquot into sealed vials.
- Incubation: Place aliquots in controlled stability chambers at 4°C (refrigerated control), 25°C (room temp), and 40°C (accelerated). Sample in triplicate at each time point (e.g., 0, 1, 3, 6 months).
- Analysis:
- Size & PDI: Measure by Dynamic Light Scattering (DLS) after gentle reconstitution/swirling.
- Drug Content: Use HPLC to quantify encapsulated vs. free drug following separation (dialysis, centrifugation).
- Morphology: Assess via TEM or SEM for structural changes.
- (Liposomes) Phospholipid Hydrolysis: Analyze via TLC or HPLC-MS for lyso-lipid formation.
Protocol 2: Shear Stress Resistance Test
- Equipment Setup: Use a rheometer with a concentric cylinder or cone-plate geometry, or a controlled syringe pump with a narrow-gauge needle/capillary.
- Shear Application: Subject nanoparticle suspension to a defined, high shear rate (e.g., 10,000 s⁻¹) for a set duration (e.g., 5-30 minutes) in the rheometer, or perform multiple extrusion cycles through a 100 nm polycarbonate membrane.
- Post-Stress Analysis: Immediately analyze the sheared sample vs. an untreated control using DLS (for size/PDI), nanoparticle tracking analysis (NTA) for concentration, and spectrophotometry for turbidity/aggregation.
Protocol 4: Freeze-Thaw & Lyophilization Stability
- Cycling: Subject suspensions to repeated freeze-thaw cycles (-80°C to 25°C).
- Lyophilization: Add cryoprotectant (e.g., 5% trehalose w/v) to suspension. Snap-freeze in liquid N2, then lyophilize for 48h.
- Reconstitution: Add water for injection, gently vortex.
- Evaluation: Measure % recovery of original size, PDI, and encapsulation efficiency. Use DSC to assess preservation of lipid Tm or polymer Tg.
Visualization: Stability Assessment Workflow
Stability Assessment Decision Workflow
The Scientist's Toolkit: Essential Research Reagents & Materials
| Item | Function in Stability Studies |
|---|---|
| PLGA (50:50) | Benchmark biodegradable polymer for NP formation; hydrolysis rate dictates degradation stability. |
| DPPC & Cholesterol | High-Tm phospholipid and stabilizing agent for forming robust, low-permeability liposomes. |
| Trehalose / Sucrose | Cryoprotectant & lyoprotectant; forms glassy matrix to protect nanostructure during freeze-drying. |
| DSPE-PEG(2000) | PEGylated lipid; provides steric stabilization, reducing aggregation in both liposomes and polymeric NPs. |
| HPLC-MS System | For quantifying drug encapsulation efficiency and detecting chemical degradation products (e.g., lyso-lipids). |
| Dynamic Light Scattering (DLS) Instrument | Critical for monitoring hydrodynamic diameter, polydispersity index (PDI), and zeta potential over time. |
| Differential Scanning Calorimeter (DSC) | Measures thermal transitions (Tg for polymers, Tm for lipids), essential for formulation optimization. |
| Polycarbonate Membranes (e.g., 100 nm) | For liposome/polymeric NP extrusion to achieve uniform size and test mechanical shear resistance. |
| Size Exclusion Columns (e.g., Sephadex G-50) | For separating encapsulated drug from free drug post-stability testing. |
This whitepaper provides a direct technical comparison of polymeric nanoparticles (PNPs) and liposomes, focusing on their pharmacokinetic (PK) profiles, biodistribution patterns, and differential utilization of the Enhanced Permeability and Retention (EPR) effect. This analysis is framed within the broader thesis that the fundamental structure and composition of these nanocarriers dictate their biological fate and therapeutic efficacy.
Core Structural & Compositional Thesis
The primary distinction lies in their core architecture:
- Polymeric Nanoparticles: Typically solid matrices (e.g., PLGA, chitosan, PEG-PLGA) where the drug is dissolved, entrapped, encapsulated, or attached. Their biodegradability, polymer crystallinity, and surface chemistry are key determinants.
- Liposomes: Phospholipid bilayer vesicles enclosing an aqueous core. Their classification (conventional, PEGylated, ligand-targeted), lipid composition, charge, and lamellarity govern their behavior.
This structural divergence directly cascades into differences in stability, drug release kinetics, protein corona formation, and interactions with biological systems.
Pharmacokinetics: Quantitative Comparison
Table 1: Comparative Pharmacokinetic Parameters of Standard 100 nm Carriers
| Parameter | Polymeric Nanoparticles (PEG-PLGA) | Liposomes (PEGylated, DPPC/Cholesterol) | Key Determinant |
|---|---|---|---|
| Circulation Half-life (t₁/₂β) | 12 - 24 hours | 24 - 48 hours | Density & integrity of PEG coating; Stealth properties |
| Volume of Distribution (Vd) | Low (0.1 - 0.5 L/kg) | Very Low (0.05 - 0.1 L/kg) | Particle rigidity & RES evasion capability |
| Clearance (CL) | Primarily MPS/Res organs | Primarily MPS/Res organs; Slower | Rate of opsonization & enzymatic degradation |
| Cmax | Dose-dependent; Often high | Dose-dependent; Sustained release profile | Injection bolus & initial RES trapping |
| AUC (Area Under Curve) | High | Typically higher than PNPs | Integrated measure of circulation longevity |
Biodistribution: Organ-Specific Accumulation
Table 2: Comparative Biodistribution (% Injected Dose per Gram of Tissue, 24h Post-IV)
| Tissue/Organ | Polymeric Nanoparticles | Liposomes (PEGylated) | Rationale & Structural Driver |
|---|---|---|---|
| Blood | Moderate | High | Liposome fluid bilayer mimics biological membranes; superior stealth. |
| Liver | High | Moderate-High | RES sequestration; influenced by size & surface hydrophobicity. |
| Spleen | High | Moderate | Filtration by splenic sinusoids; dependent on particle rigidity. |
| Tumor (via EPR) | Variable (5-10% ID/g) | Consistent (8-15% ID/g) | Liposome stability in circulation enhances EPR accumulation. |
| Kidneys | Low | Very Low | Size (>10 nm) prevents glomerular filtration. |
| Lungs | Low | Very Low | Avoidance of capillary bed entrapment. |
| Heart & Muscle | Very Low | Very Low | Lack of active targeting & poor extravasation in normal tissue. |
EPR Effect Utilization: Mechanisms and Efficacy
The EPR effect is a passive targeting mechanism relying on leaky tumor vasculature and impaired lymphatic drainage. Utilization efficacy is a direct function of nanocarrier longevity in circulation (stealth) and size (~10-200 nm).
- Polymeric Nanoparticles: Achieve EPR via sustained release and surface functionalization (e.g., PEGylation). However, batch-to-batch variability in polymer properties can lead to inconsistent tumor accumulation.
- Liposomes: The gold standard for EPR exploitation due to their tunable, well-defined phospholipid bilayers and proven ability to achieve long circulation. Their stability prevents premature drug leakage, delivering a higher payload to the tumor site.
Experimental Protocol: Evaluating EPR-Driven Tumor Accumulation
Title: Quantitative Biodistribution Study Using Radiolabeled Nanocarriers
- Nanocarrier Preparation & Radiolabeling:
- PNPs: Synthesize PEG-PLGA nanoparticles via nanoprecipitation. Radiolabel by incorporating a chelator (e.g., DOTA) into the polymer chain, followed by complexation with ¹¹¹In or using ³H-labeled polymer.
- Liposomes: Prepare PEGylated liposomes via thin-film hydration & extrusion. Radiolabel by incorporating ³H-cholesteryl hexadecyl ether (³H-CHE, a non-exchangeable, non-metabolizable lipid tracer) into the bilayer or by loading the aqueous core with ¹¹¹In via a gradient method.
- Animal Model: Implant human xenograft or murine syngeneic tumor cells subcutaneously in immunocompromised or immunocompetent mice, respectively. Allow tumors to grow to ~200-500 mm³.
- Dosing & Cohorts: Intravenously inject a known radioactive dose (e.g., 100 µCi, 1 mg nanoparticle mass) via the tail vein. Establish cohorts for multiple time points (e.g., 1, 4, 24, 48, 72 hours). Include n≥5 animals per group.
- Terminal Harvest & Analysis: At each time point, euthanize animals. Perfuse with saline via cardiac puncture. Harvest blood, tumor, liver, spleen, kidneys, lungs, heart, and a muscle sample. Weigh all tissues.
- Quantification: For ¹¹¹In, measure gamma counts per minute (CPM) in a gamma counter. For ³H, digest tissues and measure via scintillation counting. Calculate % Injected Dose per Gram (%ID/g) and % Injected Dose per Organ (%ID/organ).
- Imaging (Optional): Use SPECT/CT imaging for ¹¹¹In-labeled carriers at serial time points for longitudinal, non-invasive visualization of biodistribution.
The Scientist's Toolkit: Key Research Reagent Solutions
Table 3: Essential Materials for Comparative Nanocarrier Research
| Item/Category | Example Products/Formulations | Primary Function in Research |
|---|---|---|
| Polymer for PNPs | PLGA (RESOMER series), mPEG-PLGA (Akina), Chitosan (Sigma-Aldrich) | Forms the biodegradable, drug-encapsulating matrix of nanoparticles. |
| Phospholipids for Liposomes | DPPC, DSPC, DSPE-PEG2000 (Avanti Polar Lipids) | Building blocks of the lipid bilayer; PEG-lipids confer stealth properties. |
| Radiolabel for Tracking | ¹¹¹In-oxine, ³H-cholesteryl hexadecyl ether (PerkinElmer, American Radiolabeled Chemicals) | Enables quantitative biodistribution and PK studies via gamma/scintillation counting. |
| Size & Zeta Analyzer | Malvern Zetasizer Nano ZS | Measures hydrodynamic diameter (DLS), PDI, and surface charge (zeta potential). |
| Chromatography Media | Sepharose CL-4B, Sephadex G-50 (Cytiva) | For size-exclusion chromatography to purify nanocarriers and remove unencapsulated drug/radiolabel. |
| In Vivo Imaging System | IVIS Spectrum, Bruker SkyScan PET/CT | For non-invasive, longitudinal fluorescence or nuclear imaging of biodistribution. |
Visualizations
Diagram 1: Structural Influence on Biological Fate
Diagram 2: Experimental Workflow for EPR Quantification
The comparative analysis confirms that the foundational thesis holds: the solid, polymeric matrix of PNPs versus the aqueous-core lipid bilayer of liposomes creates distinct PK/BD profiles. Liposomes, particularly PEGylated variants, generally exhibit superior circulation longevity and more reliable EPR-mediated tumor accumulation due to their biomimetic, stable structure. PNPs offer advantages in controlled drug release and material versatility. The choice between systems for drug development hinges on the therapeutic payload, desired release kinetics, and the specific pathological vasculature targeted.
Within a thesis examining the basic structure and composition of polymeric nanoparticles (PNPs) and liposomes, a direct comparison of their biocompatibility, immunogenicity, and toxicity is paramount. The fundamental architectural differences—synthetic solid-core polymers versus lipid bilayer vesicles—dictate distinct biological interactions, determining their success in nanomedicine applications such as drug delivery, diagnostics, and vaccines.
Biocompatibility: Comparative Analysis
Biocompatibility refers to the ability of a material to perform its desired function without eliciting any undesirable local or systemic effects in the host.
Structural Determinants of Biocompatibility
- Polymeric Nanoparticles (PNPs): Typically composed of polymers like poly(lactic-co-glycolic acid) (PLGA), polyethylene glycol (PEG), or chitosan. Their biocompatibility is heavily influenced by monomer composition, molecular weight, crystallinity, and degradation rate. Acidic degradation products (e.g., from PLGA) can cause local pH drops and inflammation.
- Liposomes: Composed of natural or synthetic phospholipids (e.g., phosphatidylcholine, cholesterol). Their structure mimics biological membranes, often conferring high inherent biocompatibility. Surface charge (via cationic lipids) and lipid peroxidation potential are critical factors.
Quantitative Metrics of Biocompatibility
Table 1: Key Quantitative Biocompatibility Metrics for PNPs vs. Liposomes
| Metric | Typical Polymeric Nanoparticles (PLGA-based) | Typical Liposomes (Phosphatidylcholine/Cholesterol) | Assay/Protocol Reference |
|---|---|---|---|
| Hemolysis (% at 1 mg/mL) | 5-15% (varies with surface charge) | <2% (neutral/zwitterionic) | ISO/TR 7405-2; Incubation with RBCs, measure Hb release. |
| Protein Adsorption (Corona Thickness, nm) | 10-20 nm (dense, often irreversible) | 5-10 nm (softer, more dynamic) | DLS/SLS post-incubation with plasma; LC-MS/MS for composition. |
| Complement Activation (C3a, SC5b-9) | Moderate to High (PEGylation reduces) | Low (increased by cationic lipids) | ELISA of serum after NP exposure. |
| Degradation Half-life in vivo | Weeks to months (PLGA: Mw & ratio dependent) | Hours to days (lipid exchange, enzymatic degradation) | Radiolabel tracking, HPLC of degradation products. |
| Cell Viability (IC50 in HepG2, mg/mL) | 0.5-2.0 | >2.0 | ISO 10993-5; MTT assay after 24h exposure. |
Immunogenicity: Head-to-Head Evaluation
Immunogenicity is the potential of a nanoparticle to induce an innate or adaptive immune response, which can be detrimental (hypersensitivity, accelerated clearance) or beneficial (vaccine adjuvanticity).
Innate Immune Recognition
- PNPs: Recognized primarily via pattern recognition receptors (PRRs) for synthetic materials (e.g., scavenger receptors). PEGylated PNPs can induce anti-PEG IgM antibodies, leading to accelerated blood clearance (ABC phenomenon).
- Liposomes: Recognized by immune cells via surface opsonins or direct lipid interaction (e.g., cationic lipids activating TLRs). "Stealth" liposomes use PEG-lipids (PEG-DSPE) to reduce opsonization, but also risk anti-PEG immunity.
Key Signaling Pathways in Nanoparticle Immunogenicity
Diagram 1: Key Immune Recognition Pathways for PNPs/Liposomes
Comparative Immunogenicity Data
Table 2: Immunogenicity Profiles of PNPs vs. Liposomes
| Immune Parameter | Polymeric Nanoparticles | Liposomes | Experimental Protocol Summary |
|---|---|---|---|
| Anti-NP Antibody Induction | High for some polymers (e.g., PGA); Anti-PEG possible. | Low for conventional; high for cationic/adjuvant types. | ELISA for NP-specific IgG/IgM after i.v. boost in mice. |
| Cytokine Storm Potential | Moderate (via complement, macrophages). | High with cationic lipids (potent TLR activation). | Multiplex cytokine array (IFN-γ, IL-6, IL-1β) post-injection. |
| Accelerated Blood Clearance (ABC) | Observed with PEGylated systems. | Well-documented for PEGylated liposomes. | Radioisotope tracking of 2nd dose pharmacokinetics. |
| Dendritic Cell Maturation (%CD86+) | 30-60% (material-dependent). | 20-80% (lipid composition dependent). | Flow cytometry of bone marrow-derived DCs post 24h co-culture. |
| Inflammasome Activation (NLRP3) | Common with hydrophobic/ crystalline polymers. | Less common; associated with cholesterol crystals. | Caspase-1 assay, IL-1β ELISA in primed macrophages. |
Toxicity Profiles: Mechanisms and Metrics
Toxicity encompasses acute and chronic adverse effects, including cytotoxicity, organ-specific toxicity (e.g., hepatotoxicity, nephrotoxicity), and genotoxicity.
Primary Toxicity Mechanisms
- PNPs: Often mediated by (1) oxidative stress from catalytic surface properties, (2) lysosomal dysfunction and rupture due to slow degradation, and (3) inflammatory response to degradation products.
- Liposomes: Toxicity is frequently (1) dose-dependent, related to lipid overload in macrophages (foam cell formation), (2) reactive to cationic lipid components causing membrane disruption, and (3) related to phospholipidosis (accumulation of lipids in lysosomes).
Comparative Toxicity Data
Table 3: Systemic and Cellular Toxicity Comparison
| Toxicity Endpoint | Polymeric Nanoparticles | Liposomes | Standard Assay/Model |
|---|---|---|---|
| Maximum Tolerated Dose (MTD, mg/kg) | 50-200 (highly variable) | 100-300 (for conventional) | Rodent single-dose escalation, monitoring mortality/morbidity. |
| Hepatotoxicity (ALT/AST Elevation) | Frequent (Kupffer cell uptake, inflammation). | Lower for stealth types; high for cationic. | Serum biochemistry 24h post-i.v. administration. |
| Nephrotoxicity | Potential due to aggregation/ filtration issues. | Rare with small (<5 nm) particles. | Histopathology (glomerular injury), BUN/Creatinine. |
| Genotoxicity (Comet Assay Tail Moment) | Often positive (ROS-mediated). | Generally negative (unless loaded with genotoxic drug). | OECD 489 (In vivo alkaline comet assay). |
| Phospholipidosis Incidence | Low. | Moderate to High (lysosomal lipid accumulation). | TEM imaging of foam cells in liver/spleen; Lamellar Body assay. |
Experimental Protocols for Key Cited Assays
Protocol 1: Hemolysis Assay (ASTM E2524-08)
- Collect fresh human or animal blood in heparinized tubes.
- Isolate red blood cells (RBCs) via centrifugation (1500xg, 10 min, 4°C), wash 3x with PBS.
- Prepare a 2% (v/v) RBC suspension in PBS.
- Incubate with nanoparticle serial dilutions (0.1-2 mg/mL) for 1 hour at 37°C.
- Centrifuge, measure supernatant absorbance at 540 nm.
- Use 0.1% Triton X-100 and PBS as positive and negative controls, respectively.
- Calculate % Hemolysis = (Abssample - Absnegative)/(Abspositive - Absnegative) * 100.
Protocol 2: In Vivo Accelerated Blood Clearance (ABC) Phenomenon
- Priming Dose: Administer PEGylated nanoparticle (or control) intravenously to mice (n=5/group) at a standard dose (e.g., 5 µmol lipid/kg for liposomes).
- Resting Period: Wait 7-14 days to allow potential anti-PIgM response.
- Challenging Dose: Administer a second, identical dose containing a tracer (e.g., ³H-cholesteryl hexadecyl ether or DiR dye).
- Pharmacokinetic Sampling: Collect blood samples at multiple time points (e.g., 2 min, 30 min, 2h, 8h, 24h).
- Analysis: Quantify tracer in blood. Calculate pharmacokinetic parameters (AUC, t1/2). A significant reduction in AUC for the primed group indicates ABC.
Protocol 3: Dendritic Cell Maturation Assay
- Generate bone marrow-derived dendritic cells (BMDCs) from mice using GM-CSF (10 ng/mL) over 7 days.
- Harvest immature DCs and seed in 24-well plates (1e6 cells/well).
- Treat with nanoparticles at sub-toxic concentrations (e.g., 10-100 µg/mL) for 24 hours. Use LPS (100 ng/mL) as a positive control.
- Harvest cells, stain with fluorescent antibodies against surface markers: CD11c (DC identifier), MHC-II, CD80, CD86.
- Analyze via flow cytometry. Gate on CD11c+ cells and report the percentage of cells positive for high levels of CD86/CD80 or the mean fluorescence intensity (MFI).
The Scientist's Toolkit: Research Reagent Solutions
Table 4: Essential Materials for Comparative Nanoparticle Toxicology/Immunology Studies
| Reagent/Material | Supplier Examples | Critical Function in Experiments |
|---|---|---|
| PLGA (50:50, acid-terminated) | Sigma-Aldrich, Lactel Absorbable Polymers | The benchmark polymer for PNPs; defines degradation rate and biocompatibility. |
| 1,2-distearoyl-sn-glycero-3-phosphocholine (DSPC) | Avanti Polar Lipids, CordenPharma | The primary structural phospholipid for forming stable, rigid liposome bilayers. |
| Cholesterol (Pharma Grade) | Sigma-Aldrich, Avanti Polar Lipids | Incorporated into liposome formulations (>30 mol%) to enhance bilayer stability and reduce leakage. |
| DSPE-PEG(2000) | Avanti Polar Lipids, NOF Corporation | The gold-standard "stealth" agent for creating PEGylated (long-circulating) versions of both PNPs and liposomes. |
| Mouse Cytokine 10-Plex Panel | Thermo Fisher, Bio-Rad | Multiplex immunoassay to quantify key pro/anti-inflammatory cytokines (IL-1β, IL-6, TNFα, IFN-γ, etc.) from serum or cell media. |
| CellTiter 96 AQueous MTS Reagent | Promega | Colorimetric assay for quantifying cell viability and proliferation (lower cytotoxicity than MTT). |
| Anti-PEG IgM ELISA Kit | Alpha Diagnostic International | Specifically quantifies anti-PEG IgM antibodies in serum, critical for ABC phenomenon studies. |
| LAL Chromogenic Endotoxin Assay Kit | Lonza, Associates of Cape Cod | Essential for quantifying endotoxin levels in all nanoparticle preparations, as endotoxin is a potent confounder in immunogenicity studies. |
| ³H-Cholesteryl Hexadecyl Ether | PerkinElmer, American Radiolabeled Chemicals | Non-exchangeable, non-metabolizable radioactive tracer for accurate in vivo liposome pharmacokinetics and biodistribution studies. |
Within the ongoing thesis comparing the basic structure and composition of polymeric nanoparticles (PNPs) and liposomes, a critical empirical chapter must be dedicated to benchmarking their performance as drug delivery systems. Two of the most decisive metrics are Drug Loading Capacity (DLC) and Drug Release Kinetics. These parameters are intrinsically linked to the fundamental architecture of each system: the solid, polymeric matrix of PNPs versus the lipid bilayer(s) of liposomes. This whitepaper serves as a technical guide for conducting and interpreting these benchmark comparisons, providing standardized protocols, data presentation formats, and analytical frameworks.
Core Structural Determinants of Performance
The divergent structural foundations of PNPs and liposomes dictate their loading and release mechanisms.
- Polymeric Nanoparticles: Composed of a solid polymer matrix (e.g., PLGA, chitosan, PLA). Drug is either dispersed within the matrix (for hydrophobic drugs) or adsorbed/conjugated to the surface. DLC is governed by polymer-drug affinity, porosity, and method of preparation (nanoprecipitation, emulsion). Release is typically controlled by a combination of diffusion and polymer degradation (bulk or surface erosion).
- Liposomes: Phospholipid bilayer vesicles enclosing an aqueous core. Loading is compartment-dependent: hydrophobic drugs embed in the bilayer, while hydrophilic drugs are encapsulated in the aqueous interior. DLC depends on bilayer rigidity, lamellarity, and aqueous volume. Release is primarily governed by phospholipid permeability, membrane destabilization, or endocytosis-triggered disruption.
Experimental Protocols for Benchmarking
Protocol: Determining Drug Loading Capacity (DLC) and Encapsulation Efficiency (EE)
Objective: Quantify the amount of drug successfully incorporated into the nanocarrier.
Materials: Purified nanoparticle/liposome dispersion, model drug (e.g., Doxorubicin, Paclitaxel), centrifugation filters (MWCO 10-100 kDa), HPLC system or UV-Vis spectrophotometer.
Method:
- Prepare nanocarriers loaded with drug using a standardized method (e.g., thin-film hydration for liposomes, nanoprecipitation for PNPs).
- Separate unencapsulated/free drug from the nanoparticles using a validated method:
- Ultracentrifugation: Spin at high speed (e.g., 100,000 x g, 45 min) to pellet particles. Collect supernatant.
- Size Exclusion Chromatography (SEC): Pass dispersion through a column (e.g., Sephadex G-50) to separate particles from free drug.
- Dialysis/Membrane Filtration: Use centrifugal filters with an appropriate molecular weight cut-off.
- Lyse the purified nanoparticle fraction (using organic solvent for PNPs/lipid bilayer, or detergent for liposome aqueous core) to release all encapsulated drug.
- Quantify drug concentration in the lysate (Cloaded) and in the free-drug fraction (Cfree) using HPLC or a calibrated absorbance assay.
- Calculate key metrics:
- Encapsulation Efficiency (EE %) = [Mass of drug in nanoparticles / Total mass of drug used initially] x 100
- Drug Loading Capacity (DLC %) = [Mass of drug in nanoparticles / Total mass of drug-loaded nanoparticles] x 100
- Drug Loading (DL %) = [Mass of drug in nanoparticles / Total mass of nanoparticles] x 100
Protocol: In Vitro Drug Release Kinetics
Objective: Profile the rate and extent of drug release under simulated physiological conditions.
Materials: Dialysis bag or Float-A-Lyzer (MWCO appropriate for drug retention), release medium (e.g., PBS at pH 7.4, optionally with surfactants), shaking incubator at 37°C.
Method:
- Place a known volume of purified, drug-loaded nanocarrier dispersion into a dialysis device.
- Immerse the device in a large volume of release medium (sink conditions maintained).
- At predetermined time intervals, withdraw a sample of the external release medium and replace it with fresh medium.
- Quantify the drug concentration in each sample (Creleasedt).
- Plot the cumulative drug release (%) versus time.
- Fit release data to mathematical models to elucidate the dominant mechanism:
- Zero-order: Qt = Q0 + k_0 * t (Constant release rate)
- First-order: ln(100 - Qt) = ln(100) - k1 * t (Rate depends on remaining drug)
- Higuchi: Qt = kH * √t (Diffusion-controlled release from a matrix)
- Korsmeyer-Peppas: Qt / Q∞ = k * t^n (Power law model; 'n' indicates release mechanism)
Comparative Data Presentation
The following tables summarize typical benchmark data from recent literature, highlighting performance differences rooted in structure.
Table 1: Benchmarking Drug Loading Metrics for Common Nanocarriers
| Nanocarrier Type | Core Polymer/Lipid | Model Drug | Typical EE (%) | Typical DLC (wt%) | Key Structural Factor Influencing Load |
|---|---|---|---|---|---|
| Liposome | HSPC/Chol/DSPE-PEG | Doxorubicin (Hydrophilic) | 85 - 98 | 5 - 15 | Aqueous core volume, active loading (pH gradient) |
| Liposome | DPPC/Chol | Paclitaxel (Hydrophobic) | >90 | 1 - 5 | Bilayer composition and fluidity |
| Polymeric NP (PLGA) | PLGA-PEG | Doxorubicin (Hydrophilic) | 50 - 80 | 3 - 10 | Drug-polymer interaction, porosity |
| Polymeric NP (PLGA) | PLGA | Paclitaxel (Hydrophobic) | 70 - 95 | 5 - 25 | Hydrophobic partitioning into matrix |
Table 2: Comparative Drug Release Kinetics (Model Fitting Parameters)
| Nanocarrier Type | Drug | Dominant Release Model (PBS, 37°C) | Model Parameter (n or k) | Approx. Time for 80% Release (T_80%) | Implied Release Mechanism |
|---|---|---|---|---|---|
| Liposome | Doxorubicin | First-order | k_1 ≈ 0.05 h⁻¹ | 24 - 48 h | Permeability-controlled leakage |
| Liposome (Thermosensitive) | Doxorubicin | Burst release, then first-order | n/a | <1 h (at Tm) | Phase transition-induced rupture |
| Polymeric NP (PLGA) | Paclitaxel | Higuchi / Korsmeyer-Peppas | n ≈ 0.45 | 7 - 28 days | Diffusion-controlled, coupled with erosion |
| Polymeric NP (Chitosan) | siRNA | Burst release, then plateau | n/a | 2 - 8 h | Surface desorption / rapid dissociation |
Visualization of Workflows and Relationships
Title: DLC & Release Benchmarking Workflow
Title: Structure Determines Loading & Release
The Scientist's Toolkit: Essential Research Reagents & Materials
Table 3: Key Reagent Solutions for Benchmarking Experiments
| Item/Category | Example Products & Specifications | Primary Function in Benchmarking |
|---|---|---|
| Model Drugs | Doxorubicin HCl (hydrophilic), Paclitaxel (hydrophobic), Fluorescent dyes (DiI, Coumarin-6). | Serve as payloads to quantify loading efficiency and track release. Different solubilities test carrier versatility. |
| Polymer for PNPs | PLGA (50:50, varied MW), PLGA-PEG, Chitosan, Polycaprolactone (PCL). | Forms the solid matrix of nanoparticles. MW, crystallinity, and end-group determine degradation rate and drug affinity. |
| Lipids for Liposomes | HSPC, DPPC, Cholesterol, DSPE-PEG2000, DOTAP. | Form the bilayer structure. Lipid chain length/saturation defines phase transition temperature (Tm) and permeability. |
| Purification Devices | Sephadex G-50 Columns, Amicon Ultra Centrifugal Filters (e.g., 100 kDa MWCO). | Separate unencapsulated free drug from the formulated nanocarriers for accurate EE/DLC measurement. |
| Release Study Apparatus | Dialysis Tubing (SnakeSkin), Float-A-Lyzer devices (various MWCO). | Provide a semi-permeable barrier to contain nanoparticles while allowing released drug to diffuse into sink medium. |
| Quantification Instruments | HPLC with C18 column & UV/FLD detector, UV-Vis Spectrophotometer, Fluorescence Plate Reader. | Precisely measure drug concentration in complex samples (lysates, release media) for calculating EE, DLC, and release profiles. |
| Buffer & Media | Phosphate Buffered Saline (PBS, pH 7.4), HEPES Buffer, Simulated Biological Fluids (e.g., SBF). | Provide controlled ionic strength and pH for stability and release studies, simulating physiological conditions. |
The development of nanomedicines, primarily polymeric nanoparticles and liposomes, represents a cornerstone of modern drug delivery. This whitepaper examines the clinical translation success of these platforms, framed within a thesis on their fundamental structural and compositional differences. While liposomes are lipid-bilayer vesicles offering high biocompatibility and efficient encapsulation of hydrophilic drugs, polymeric nanoparticles—composed of synthetic or natural polymers—provide superior structural integrity, controlled release kinetics, and versatility in surface functionalization. These core material properties directly dictate pharmacological behavior, manufacturing scalability, and ultimately, clinical and commercial outcomes. This document provides a technical analysis of approved nanotherapeutics and active clinical pipelines, linking their success to the intrinsic advantages and limitations of their underlying platform technology.
Approved Nanotherapeutics: A Platform-Based Analysis
The clinical approval of nanomedicines has been historically dominated by liposomal formulations, owing to their earlier development and biomimetic structure. Polymeric nanoparticles have gained significant traction, particularly in oncology and vaccine delivery. The following table summarizes key approved agents, categorized by platform.
Table 1: Selected Approved Nanotherapeutics (2020-2024)
| Product Name (Generic) | Platform Type | Indication (Approval Year) | Core Composition & Key Structural Feature |
|---|---|---|---|
| Onpattro (patisiran) | Lipid Nanoparticle (LNP) | hATTR Amyloidosis (2018) | Ionizable lipid, PEG-lipid, cholesterol, phospholipid. siRNA delivery. |
| Comirnaty (BNT162b2) | LNP | COVID-19 Prevention (2020/21) | Ionizable lipid (ALC-0315), PEG-lipid, cholesterol, DSPC. mRNA delivery. |
| Spikevax (mRNA-1273) | LNP | COVID-19 Prevention (2020/22) | Ionizable lipid (SM-102), PEG-lipid, cholesterol, DSPC. mRNA delivery. |
| Vyxeos (daunorubicin/cytarabine) | Liposome | AML (2017) | "CPX-351" liposome with fixed 5:1 molar ratio of drugs to lipids. |
| Onivyde (irinotecan) | Liposome | Pancreatic Cancer (2015) | PEGylated liposome encapsulating irinotecan succinate. |
| Abraxane (nab-paclitaxel) | Protein-bound Nanoparticle | Breast Cancer (2005) | Albumin-bound paclitaxel (130 nm). A polymeric protein particle. |
| Genexol-PM | Polymeric Micelle | Breast Cancer (Korea, 2007) | PEG-PLA block copolymer micelle encapsulating paclitaxel. |
| COVID-19 Vaccine (Valneva) | Virosome | COVID-19 Prevention (2022) | Influenza virus-derived membrane vesicle containing spike protein. |
Current Clinical Trial Pipeline Analysis
The pipeline reflects a shift towards advanced functionalities. Polymeric nanoparticles are increasingly used for targeted delivery and combination therapies, while next-generation liposomes and LNPs focus on nucleic acid delivery and triggered release.
Table 2: Representative Active Clinical-Stage Nanomedicines (Phase II/III)
| Candidate / Identifier | Platform | Phase | Indication | Key Differentiator |
|---|---|---|---|---|
| BNT162b2 (Omicron adapted) | LNP | III | COVID-19 | Updated mRNA payload. |
| mRNA-1647 & mRNA-1653 | LNP | II | CMV, hMPV/PIV3 | Multivalent mRNA vaccine candidates. |
| ARCT-154 (VNN-001) | LNP | III | COVID-19 | Self-amplifying mRNA. |
| CRLX-101 | Cyclodextrin-based Polymer Nanoparticle | II | Oncology | Camptothecin conjugate; PEGylated, ~20-40 nm. |
| NU-0129 (Spherical Nucleic Acid) | Gold Nanoparticle Core | I | Glioblastoma | siRNA conjugated to gold nanoparticle core. |
| LiPlaCis | Liposome | II | Metastatic Breast Cancer | Triggered by secretory PLA2 in tumor microenvironment. |
| ThermoDox | Lyso-thermosensitive Liposome | III | Hepatocellular Carcinoma | Heat-triggered release (with RFA). |
Experimental Protocols for Critical Characterization
Protocol 1: Determination of Encapsulation Efficiency (EE%) and Drug Loading (DL%) for Polymeric Nanoparticles (e.g., PLGA NPs)
- Preparation: Synthesize drug-loaded nanoparticles via single/double emulsion or nanoprecipitation.
- Separation: Isolate nanoparticles from free, unencapsulated drug via ultracentrifugation (e.g., 40,000 rpm, 45 min, 4°C) or size exclusion chromatography.
- Quantification:
- Direct Method (for EE%): Lyse the purified nanoparticle pellet in a suitable solvent (e.g., acetonitrile for PLGA). Quantify the total drug amount (Wloaded) via HPLC/UV-Vis against a standard curve.
- Indirect Method (for EE%): Quantify the amount of free drug (Wfree) in the supernatant after centrifugation.
- Calculate EE% = (Wloaded / Winitial) x 100 or EE% = [(Winitial - Wfree) / Winitial] x 100.
- Calculate DL% = (Weight of loaded drug / Total weight of nanoparticle) x 100.
- Key Parameters: Centrifugal force/time, lysing efficiency, analytical method sensitivity.
Protocol 2: In Vitro Release Kinetics Study under Sink Conditions
- Setup: Place a known amount of purified, drug-loaded nanocarrier (liposome or polymeric NP) into a dialysis bag (MWCO ~10-14 kDa) or a Franz diffusion cell donor compartment.
- Release Media: Immerse in a large volume (>10x the saturation volume) of PBS (pH 7.4) at 37°C, with gentle agitation. For simulating endosomal conditions, include a secondary study at pH 5.5. Addition of 0.1-1% w/v Tween 80 can maintain sink conditions for hydrophobic drugs.
- Sampling: At predetermined intervals, withdraw and replace an aliquot of the release medium.
- Analysis: Quantify drug concentration in aliquots using HPLC or UV-Vis spectroscopy.
- Modeling: Plot cumulative release (%) vs. time. Fit data to kinetic models (e.g., zero-order, first-order, Higuchi, Korsmeyer-Peppas) to elucidate release mechanisms.
Protocol 3: Evaluation of Cellular Uptake Mechanism via Pharmacological Inhibition
- Cell Seeding: Plate adherent cells (e.g., HeLa, MCF-7) at ~70% confluency in 24-well plates.
- Pre-treatment: Incubate cells for 30-60 min with specific endocytic inhibitors: Chlorpromazine (10 µg/mL) for clathrin-mediated endocytosis; Genistein (200 µM) for caveolae-mediated endocytosis; Amiloride (1 mM) for macropinocytosis; Sodium Azide (10 mM) + 2-Deoxyglucose (50 mM) for energy depletion (4°C).
- Nanoparticle Exposure: Add fluorescently-labeled nanoparticles (liposomes or polymeric NPs) to wells (with inhibitor still present) for 2-4 hours.
- Analysis: Wash cells thoroughly, trypsinize, and analyze mean fluorescence intensity (MFI) via flow cytometry. Compare MFI of inhibitor-treated groups to untreated control.
- Interpretation: >50% reduction in MFI relative to control indicates the inhibited pathway is a major uptake route.
Visualizing the Endocytic Pathways for Nanocarrier Internalization
Title: Nanocarrier Cellular Uptake and Intracellular Trafficking Pathways
The Scientist's Toolkit: Key Reagent Solutions for Nanocarrier Development
Table 3: Essential Research Reagents and Materials
| Reagent / Material | Function & Explanation |
|---|---|
| 1,2-distearoyl-sn-glycero-3-phosphocholine (DSPC) | A saturated, high-phase-transition phospholipid. Provides structural rigidity and low permeability to liposome membranes, enhancing blood circulation time. |
| DSPE-PEG2000 | Phospholipid conjugated to polyethylene glycol (PEG). The PEG chain creates a steric barrier ("stealth" effect) around the liposome/nanoparticle, reducing opsonization and clearance by the mononuclear phagocyte system. |
| Ionizable Lipid (e.g., DLin-MC3-DMA, ALC-0315) | A key component of LNPs. Remains neutral at physiological pH but acquires a positive charge in acidic endosomes, promoting membrane disruption and cytoplasmic release of nucleic acid payloads (proton sponge/fusion). |
| PLGA (Poly(lactic-co-glycolic acid)) | A biodegradable, FDA-approved copolymer. The workhorse polymer for nanoparticle formation. Degradation rate and drug release kinetics can be tuned by altering the lactic:glycolic acid ratio and molecular weight. |
| mPEG-PLA Diblock Copolymer | Forms polymeric micelles. The hydrophobic PLA core encapsulates poorly soluble drugs, while the hydrophilic PEG corona provides solubility and stealth properties. Used in products like Genexol-PM. |
| Dialysis Tubing (MWCO 3.5-14 kDa) | Used for purifying nanoparticles from free solutes (e.g., drugs, surfactants) and for conducting in vitro release studies by allowing diffusion of small molecules while retaining nanoparticles. |
| Dynamic Light Scattering (DLS) / Zetasizer | Instrument suite for critical quality attributes: measures hydrodynamic diameter (size), polydispersity index (PDI, size distribution), and zeta potential (surface charge) of nanocarriers in suspension. |
| Size Exclusion Chromatography (SEC) Media (e.g., Sepharose CL-4B) | Gel filtration media for gentle, non-destructive purification of nanoparticles from unencapsulated materials, based on differences in hydrodynamic volume. |
The selection of an appropriate nanocarrier is a pivotal early-stage decision in nanomedicine development. This choice must be grounded in a deep understanding of the basic structure and composition of the two leading platforms: liposomes and polymeric nanoparticles (PNPs). Liposomes are spherical vesicles composed of one or more concentric phospholipid bilayers surrounding an aqueous core, mimicking biological membranes. Polymeric nanoparticles are typically solid colloidal particles, where the drug is dissolved, entrapped, encapsulated, or attached to a polymer matrix. The core thesis guiding this framework is that the optimal carrier emerges from aligning the drug's physicochemical properties and therapeutic goals with the inherent structural advantages and limitations of each system.
Core Structural & Compositional Comparison
The fundamental architecture dictates function. The table below summarizes key quantitative and qualitative distinctions.
Table 1: Fundamental Characteristics of Liposomes vs. Polymeric Nanoparticles
| Characteristic | Liposomes | Polymeric Nanoparticles (PNPs) |
|---|---|---|
| Core Composition | Aqueous interior (hydrophilic core) | Solid or matrix-like polymer core (hydrophobic/hydrophilic) |
| Shell/Matrix Material | Phospholipid bilayers (e.g., DSPC, DOPC, cholesterol) | Biodegradable polymers (e.g., PLGA, PLA, chitosan, alginate) |
| Typical Size Range | 50 – 1000 nm | 10 – 1000 nm |
| Drug Loading | Entrapment Efficiency: Highly variable (5-50%+). Hydrophilic drugs in core; hydrophobic in bilayer. | Encapsulation Efficiency: Often higher for hydrophobic drugs (30-90%+). Can be tailored via polymer-drug affinity. |
| Structural Integrity | Dynamic, can fuse, aggregate. Membrane fluidity tunable. | Generally more rigid and stable colloidal structure. |
| Surface Modification | Straightforward; PEG-lipids inserted into bilayer. | Requires chemical conjugation or coating adsorption. |
| Scalability & Manufacturing | Established (thin-film hydration, extrusion). Scale-up can be complex. | Versatile (nanoprecipitation, emulsion methods). Often considered more scalable. |
| Regulatory Precedent | Extensive (Doxil, Onivyde, mRNA vaccines). | Growing, but fewer approved products (e.g., Genexol-PM). |
| Primary Drug Compatibility | Ideal for: Hydrophilic (aqueous core), amphiphilic (bilayer), and nucleic acids. | Ideal for: Hydrophobic/ lipophilic drugs (within matrix), proteins/peptides (with specific polymers). |
Key Decision Factors & Experimental Validation Protocols
Factor 1: Drug Physicochemical Properties
The drug's Log P, solubility, and stability are primary drivers.
Protocol 1.1: Determining Optimal Carrier via Partition Coefficient Analysis
- Objective: To experimentally determine whether a drug candidate preferentially partitions into a lipid bilayer or a polymeric matrix.
- Materials:
- Test Compound: Drug candidate.
- Lipid Film: Prepared from phosphatidylcholine/cholesterol (7:3 molar ratio).
- Polymer Solution: PLGA in acetone (e.g., 10 mg/mL).
- Buffer: PBS, pH 7.4.
- Separation Devices: Ultrafiltration centrifugal units (100 kDa MWCO) or mini-columns.
- Method:
- Liposome System: Hydrate the lipid film with a drug-containing PBS solution. Extrude through 100 nm polycarbonate membranes. Separate unencapsulated drug using size-exclusion chromatography or centrifugation.
- PNP System: Formulate PNPs via nanoprecipitation—add polymer solution in acetone to an aqueous phase containing the drug under stirring. Purify by centrifugation.
- Analysis: Measure drug concentration in the purified nanocarrier fraction and the free fraction via HPLC/UV. Calculate encapsulation efficiency (EE%) and drug loading (DL%).
- Decision Point: Higher EE% in one system provides a strong initial indicator of compatibility.
Factor 2: Release Kinetics Profile
Desired release profile (sustained, triggered, burst) is contingent on carrier structure.
Protocol 2.1: In Vitro Drug Release Kinetics (Dialysis Method)
- Objective: To characterize and compare the release profile of the drug from liposomal vs. PNP formulations.
- Materials:
- Formulations: Liposome and PNP batches loaded with drug.
- Release Medium: PBS (pH 7.4) or biorelevant medium, with or without 0.1-1% w/v surfactant (e.g., Tween 80) to maintain sink conditions.
- Dialysis Device: Float-A-Lyzer or Slide-A-Lyzer cassettes (appropriate MWCO relative to drug and carrier).
- Method:
- Place a known volume of nanocarrier suspension into the dialysis device.
- Immerse the device in a large volume of release medium under gentle agitation at 37°C.
- At predetermined time points, sample the external medium and replace with fresh medium.
- Quantify released drug via HPLC.
- Decision Point: PNPs (especially from polymers like PLGA) typically exhibit slower, more sustained release due to polymer erosion/diffusion. Liposomes often show faster, diffusion-controlled release, which can be modulated with cholesterol or triggered lipids.
Factor 3: Biological Fate & Targeting Strategy
The journey from administration to target site is visualized in the following pathway.
Diagram Title: Biological Pathway of Systemically Administered Nanocarriers.
Protocol 3.1: Assessing Stealth Properties & Macrophage Uptake
- Objective: To compare the protein adsorption and macrophage association of liposome vs. PNP formulations.
- Materials:
- Formulations: PEGylated and non-PEGylated versions of both liposomes and PNPs.
- Protein Source: Fetal Bovine Serum (FBS) or human plasma.
- Cell Line: RAW 264.7 murine macrophage cells.
- Label: Fluorescent dye (e.g., DiI, Cy5) for particle tagging.
- Method:
- Protein Corona: Incubate nanoparticles with 50% FBS for 1h at 37°C. Isolate particles by centrifugation. Analyze bound proteins via SDS-PAGE or LC-MS/MS.
- Cell Uptake: Seed macrophages in 24-well plates. Treat with fluorescent nanoparticles (equivalent surface charge/size) for 2-4h. Wash, trypsinize, and analyze mean fluorescence intensity via flow cytometry.
- Decision Point: Liposomes offer more straightforward PEG incorporation for stealth. PNPs may have more variable corona profiles. Lower macrophage uptake indicates longer circulation potential.
The Scientist's Toolkit: Essential Research Reagents & Materials
Table 2: Key Reagent Solutions for Formulation & Characterization
| Reagent / Material | Primary Function | Example in Liposomes | Example in PNPs |
|---|---|---|---|
| Phospholipids (DSPC, DOPC) | Forms the structural bilayer matrix. Determines membrane fluidity and stability. | DSPC for rigid, high-Tm bilayers; DOPC for fluid bilayers. | Not applicable. |
| Cholesterol | Modulates membrane fluidity and integrity; reduces drug leakage. | Standard component (30-50 mol%). | Not typically used. |
| PEG-lipid (e.g., DSPE-PEG2000) | Provides steric stabilization ("stealth") to reduce MPS clearance. | Post-insertion or co-formulated. | PEG-PLGA diblock copolymers are used. |
| Biodegradable Polymer (PLGA) | Forms the solid, drug-encapsulating matrix. Degradation rate controls drug release. | Not applicable. | PLGA 50:50 (standard), varying MW for different release kinetics. |
| Stabilizer/Surfactant (PVA, Poloxamer) | Prevents aggregation during formulation. | May be used in some methods. | Critical in emulsion methods (e.g., PVA in single/double emulsion). |
| Size Exclusion Chromatography Media (Sephadex G-50, Sepharose CL-4B) | Purifies nanoparticles from unencapsulated drug and small molecules. | Standard for liposome purification. | Used for certain PNP types (e.g., nanogels). |
| Dialysis Membrane/Tubing | Purifies nanoparticles or conducts in vitro release studies. | Used for purification or release. | Standard for in vitro release studies. |
| Fluorescent Lipophilic Tracer (DiI, DiD) | Labels the lipid phase for imaging and cellular uptake studies. | Directly integrates into bilayer. | Can integrate into polymeric matrix if sufficiently lipophilic. |
Integrated Decision Workflow
The following logic diagram synthesizes the key decision factors into a recommended workflow.
Diagram Title: Decision Logic for Carrier Selection.
The decision between liposomes and polymeric nanoparticles is not arbitrary but a direct consequence of their foundational structure and composition. Liposomes, with their aqueous core and biomimetic bilayer, excel at delivering hydrophilic agents, amphiphiles, and genetic material, often with more straightforward surface engineering. Polymeric nanoparticles, with their solid polymer matrix, offer superior control over sustained release profiles and high loading for hydrophobic drugs, potentially with advantageous scalability. This framework advocates for a systematic, factor-driven experimental approach, beginning with partition and release studies, to align the drug candidate's profile with the inherent strengths of each nanocarrier architecture, thereby de-risking the development pathway.
Conclusion
Polymeric nanoparticles and liposomes represent distinct yet complementary paradigms in nanomedicine, each with unique structural advantages and inherent limitations. Liposomes excel in encapsulating hydrophilic drugs and have a proven clinical track record, while polymeric nanoparticles offer superior stability, controlled release kinetics, and versatility in polymer selection. The optimal choice is not universal but depends critically on the drug's physicochemical properties, the desired release profile, the route of administration, and the target pathology. Future directions point toward hybrid systems, stimuli-responsive materials, and increasingly sophisticated targeting strategies. As characterization and manufacturing techniques advance, the rational design of these nanocarriers will continue to drive innovation in personalized and precision therapeutics, solidifying their indispensable role in the next generation of drug delivery systems.